News
Subscribe to the Australian BioCommons monthly newsletter or read previous editions
Key molecular structure prediction datasets now available in the new Structural Biology AI Reference Collection
The prediction and analysis of protein structure by Australian researchers has been accelerated by the publication of the Structural Biology AI Reference Collection at the National Computational Infrastructure (NCI).
The prediction and analysis of protein structure by Australian researchers has been accelerated by the publication of the Structural Biology AI Reference Collection at the National Computational Infrastructure (NCI). This collection includes key datasets required to support deep learning models for molecular structure prediction. Its creation is a result of a targeted collaboration between Australian BioCommons, UNSW Sydney and NCI staff.
How does this new resource help researchers?
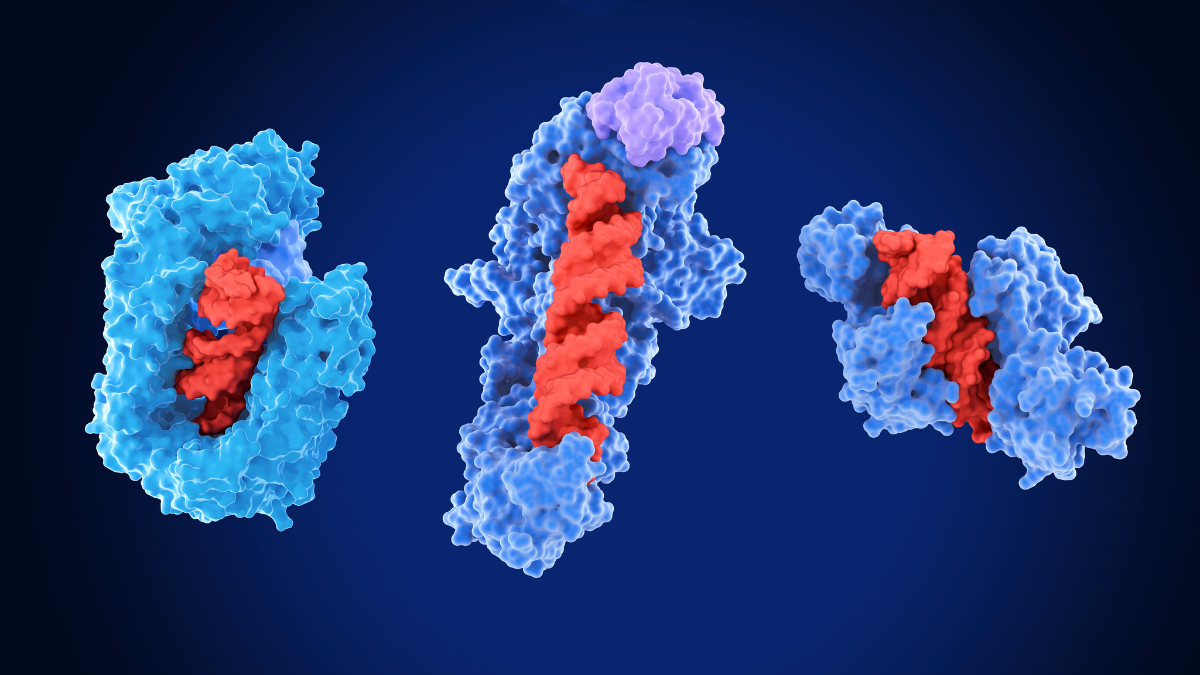
The Structural Biology AI Reference Collection includes several replicated datasets required to support protein structure research, including various deep learning models for molecular structure prediction. These copies of sequence and structure databases include UniProt, MGnify and the PDB will be regularly updated, providing a citable, versioned resource and ensuring reproducible research. The datasets support higher quality model predictions using tools such as AlphaFold3, AlphaFold2, Boltz, RoseTTaFold-All-Atom and HelixFold3.
3D molecular visualisations showing the complex structural interactions between proteins and DNA (Image: Science Photo Library)
Created through national collaboration
A team from UNSW comprising Dr Tom Litfin (Australian BioCommons) and Joshua Caley (Structural Biology Facility) worked directly with the NCI Data Collections team to add the new resources to the NCI Data Catalogue. NCI is an important partner in the BioCommons BioCLI and Workflow Commons projects, and this data collection is an example of our shared ambitions to streamline processing and analysis of molecular data at scale, and establish national ecosystems that support life science researchers at the community level.
How can researchers access the data?
The Structural Biology AI Reference Collection is now freely available. Existing NCI users can register for file-system access through the MyNCI User Portal. New users should review the information on the data products, licences, and data access to get started.
The Structural Biology AI Reference Collection is supported by the Australian Structural Biology Computing community, Australian BioCommons, and the UNSW Structural Biology Facility.
Meet the Team: Mok, UX Designer
Our team members bring deep expertise and broad experience. Hear how a UX Designer contributes to the BioCommons mission from Mok.
Describe your role at BioCommons
I’m a user experience (UX) Designer at BioCommons, which means that I help improve infrastructure and scientific research by being the friendly translator between complex systems and real humans. This type of role is new in the life sciences field, making it a lot of fun, as I sit at the intersection of science, data, and human-centred design, helping researchers, bioinformaticians, and software engineers make sense of the inherently complex biological tools and platforms they use everyday. The goal is to make it look easy - even when it’s not!
It’s challenging work, because the problems are big and complex. You’re designing for expert users, emerging technologies, and systems that genuinely matter.
There’s a lot to learn, a lot to ask, and plenty of moments where curiosity and collaboration are essential. This makes the role deeply rewarding, as your work doesn’t just improve usability; it helps accelerate research, supports discovery, and amplifies the impact of national life-science infrastructure.
How can UX Designers improve infrastructure and/or scientific research?
Think of scientific infrastructure as a powerful machine: data platforms, tools, pipelines, and services that can do amazing things. A UX Designer ensures that people can actually use that power without needing a PhD in ‘figuring stuff out’.
By taking complicated processes and reshaping them into smooth, logical journeys, we turn confusion into clarity. This saves researchers time and frustration - when tools are intuitive, scientists spend less time wrestling with interfaces and more time doing what they love: discovering, analysing, and innovating.
Good UX also makes infrastructure more accessible. It opens the door for students and early-career researchers to use advanced systems confidently, rather than those tools being limited to the experts that already know the ropes. By asking the right questions early and understanding user needs upfront, we help teams build the right thing the first time. This reduces rework and ensures that research workflows flow smoothly, helping insights travel from idea to impact more quickly.
What is the real-world impact of human-centred design at BioCommons?
At BioCommons, my impact is all about making powerful research infrastructure feel simple, friendly, and usable. I focus on turning complex scientific tools into clear experiences, helping researchers spend less time navigating systems and more time doing great science.
By listening to users and smoothing out workflows, I help ensure that BioCommons tools are not just functional, but adopted and used to their full potential. In short: I make hard things easier, science faster, and national infrastructure more human.
What makes solving scientific problems so rewarding?
This is not your average UX gig, and that’s exactly the point. I get to work closely with scientists, engineers, product leads, and stakeholders who are passionate about what they do and who will happily stretch my thinking.
It is incredibly rewarding to work in a space where UX isn’t just ‘nice to have’, but genuinely transformative. There is a unique joy in those moments where a complex process suddenly becomes clear and usable.
In short: it’s a role for UX designers who like their work meaningful, their challenges meaty, and their wins shared with science itself!
Strengthening Australia’s Nextflow community through local and global collaboration
The Sydney satellite of the international nf-core Hackathon brought together members of the Nextflow and nf-core community to work collaboratively on a diverse range of projects.
The Australian satellite site of the 2026 nf-core Hackathon has just taken place at the BioCommons node at Sydney Informatics Hub, the University of Sydney. The hands-on event brought together members of Australia’s Nextflow and nf-core community to contribute to nf-core projects, while strengthening connections locally and across the global community.
Twelve participants from University of NSW, University of Sydney, and University of Melbourne came together to collaborate in person on cutting-edge nf-core projects, with support from two Nextflow Ambassadors on BioCommons’ team, Dr Georgie Samaha and Dr Ziad Al Bkhetan.
As one of 29 global hackathon sites, the Australian team contributed to the international nf-core effort through asynchronous collaboration. Given our geography, the team was among the first to kick off the work, closing each day by connecting with the APAC teams working from sites in Aotearoa New Zealand and South Korea.
Attendees at the satellite site at the Sydney Informatics Hub, University of Sydney
The group made meaningful contributions to several nf-core projects, working to: close open issues and make a release for the nf-core ProteinFold pipeline; develop a new ConfigBuilder tool in the nf-core tools suite that helps users build Nextflow configuration files; and edit Nextflow for HPC training materials to support implementation by others with new documentation for trainers.
It was wonderful to see the progress made by the productive participants over three days together, and the outcomes reflect the depth of expertise and collaborative spirit of the Australian community. We are grateful to all participants for the energy and enthusiasm they brought. Thanks also to the Seqera team and the nf-core community for their support for the Australian satellite site of the hackathon, and especially for sponsoring our delicious lunch together on the final day!
Want to learn how to run and customise nf-core workflows? The Unlocking nf-core: customising workflows for your research workshop is taking applications now, and registrations are also open for our upcoming webinar Using Containers in Nextflow.
AI for Science Hackathon connects industry with research to accelerate Australian science
Australian BioCommons partnered with NVIDIA, OpenACC, Monash University, NCI, Pawsey and SHARON AI to bridge the gap between industry hardware experts and life science researchers.
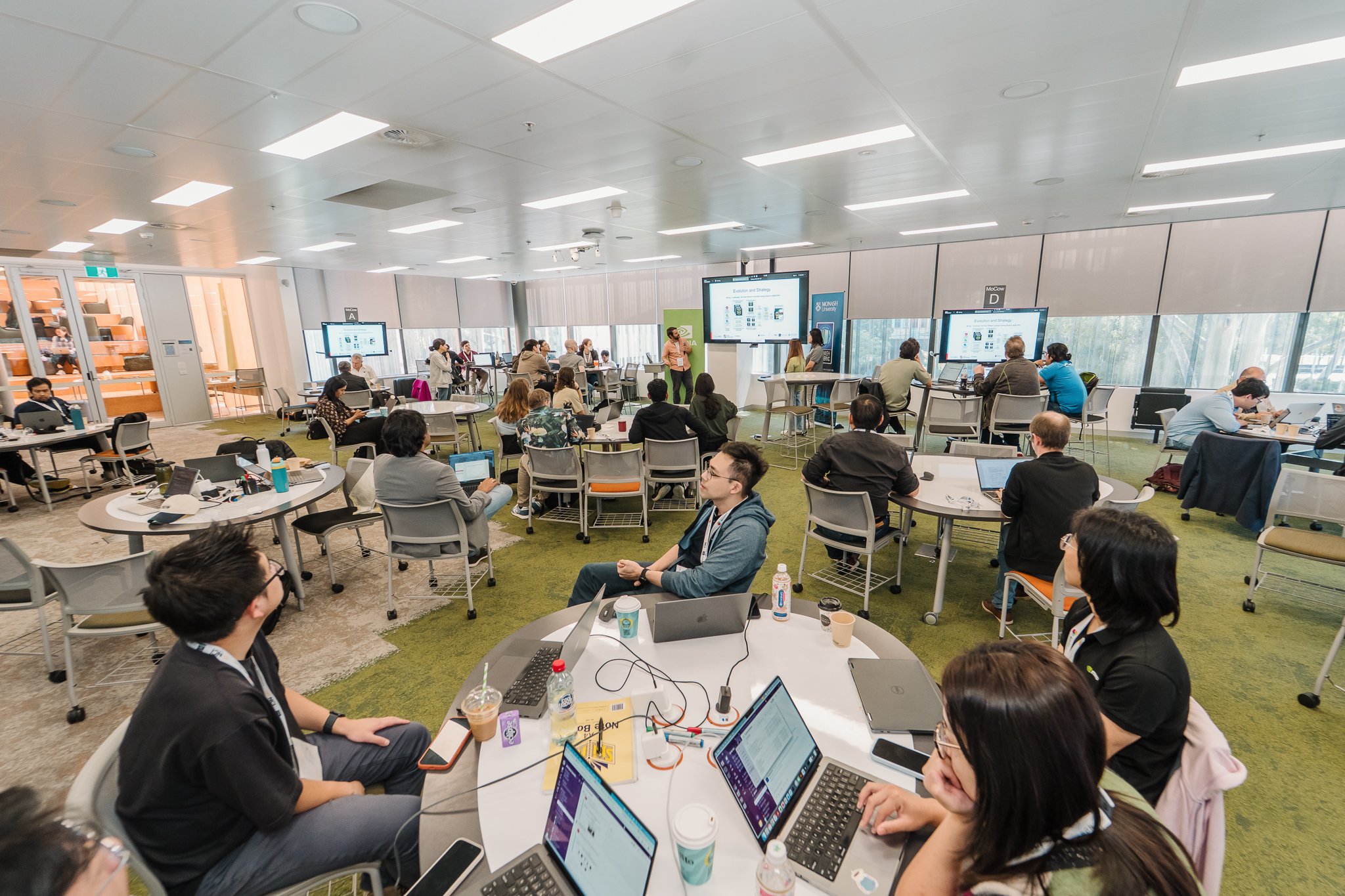
The AI for Science Australian Hackathon recently wrapped up in Melbourne, bringing together researchers and technical experts to port, accelerate and optimise research applications. The week-long event paired a select group of research projects with computational mentors and powerful compute tailored to their needs. Organised in collaboration with NVIDIA and OpenACC and hosted at Monash University, BioCommons also joined Pawsey Supercomputing Research Centre, National Computational Infrastructure and SHARON AI to boost projects across structural biology, advanced engineering, quantum chemistry and climate modelling.
What life science challenges were tackled?
Several teams leveraged the opportunity to translate software and hardware optimisations to existing medical and human health challenges. A key factor for teams applying to the event was to bring a mature project that could utilise the available mentorship and compute resources effectively.
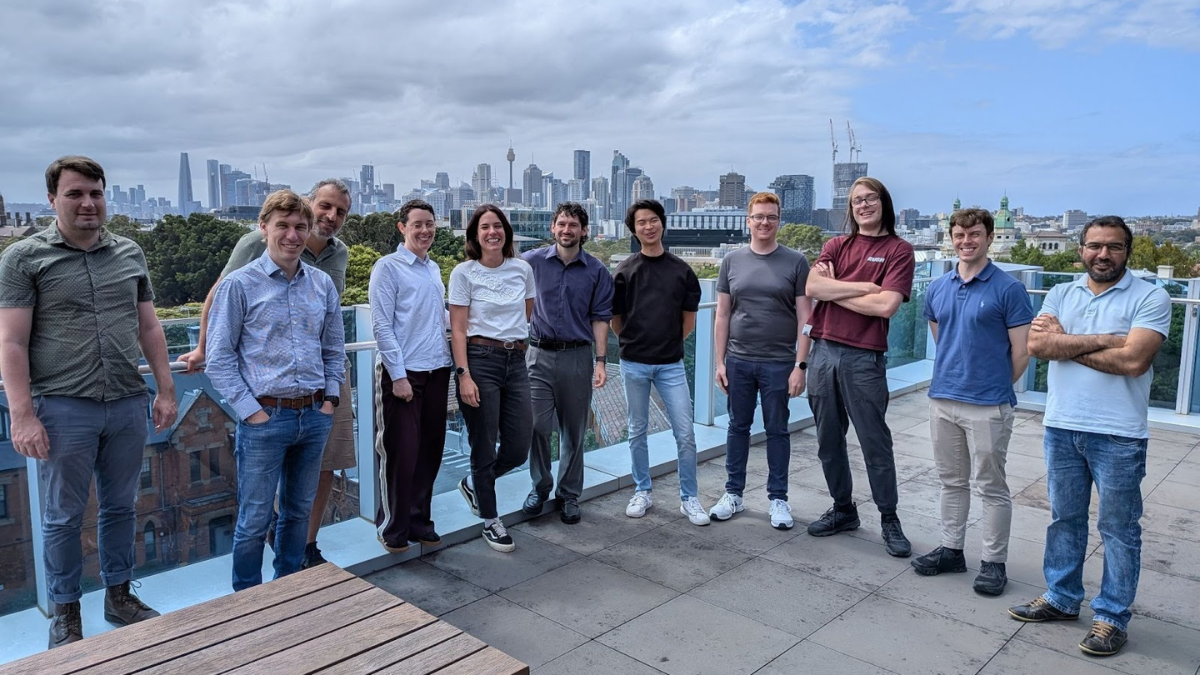
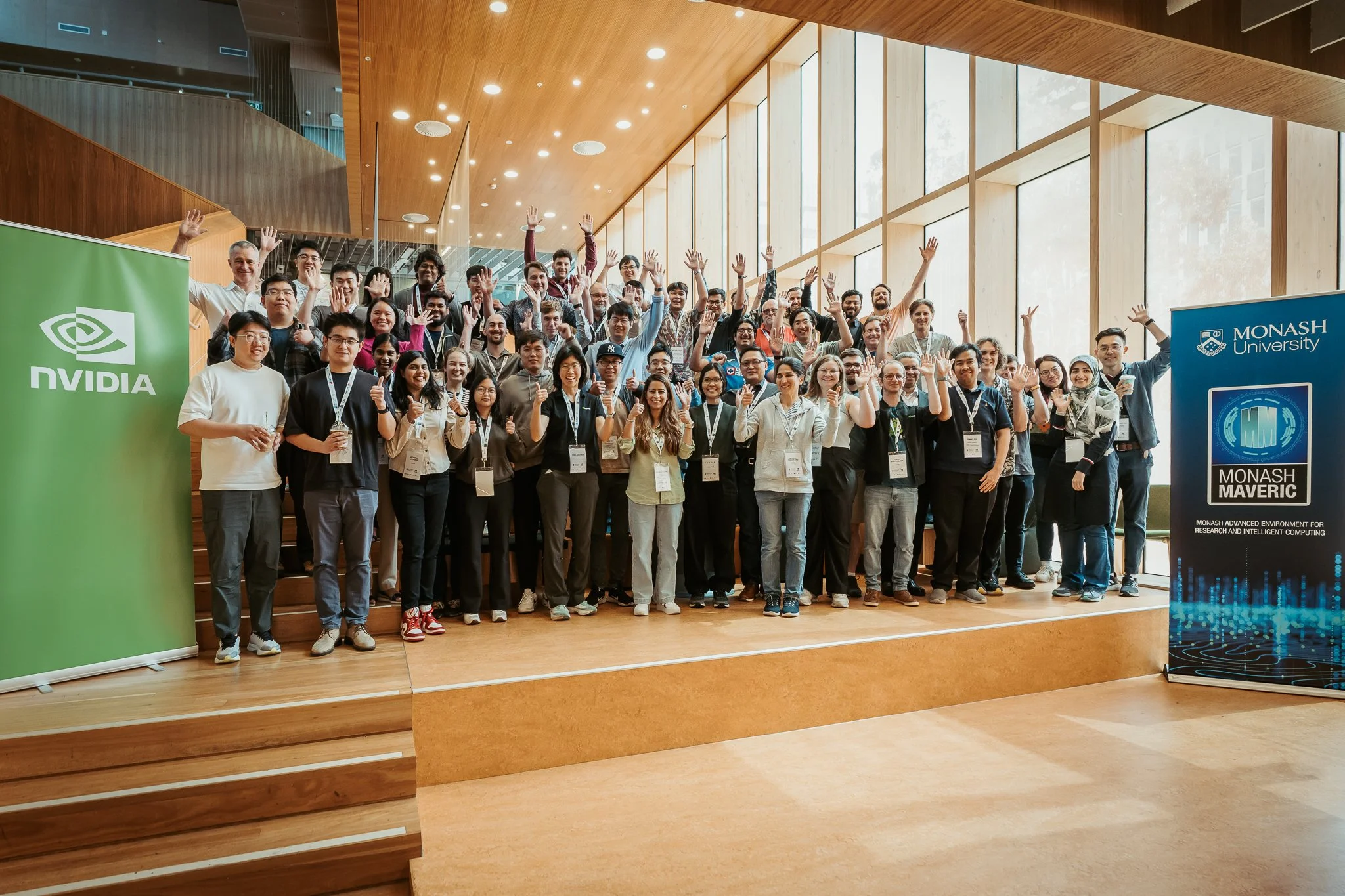
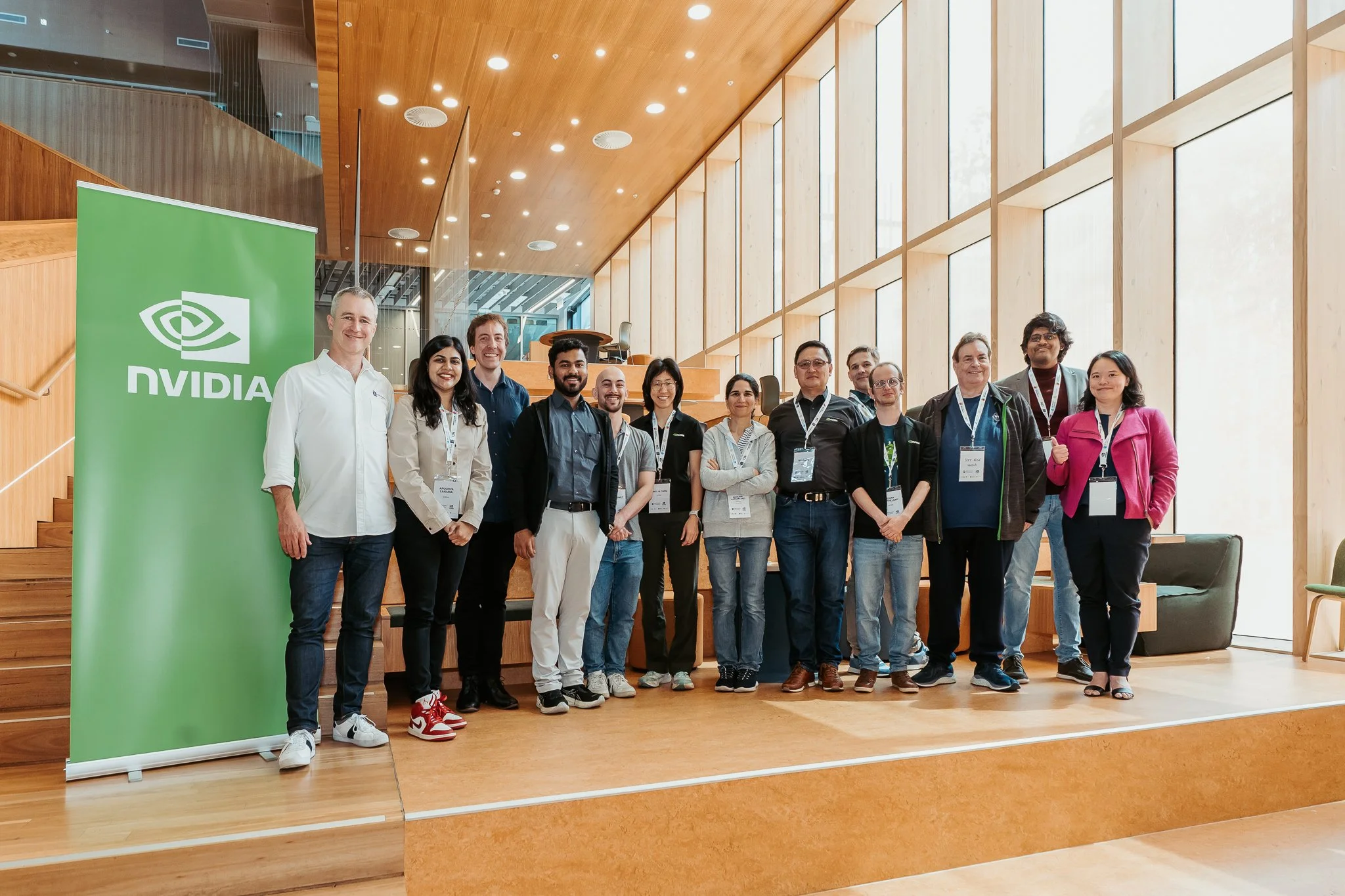
Attendees, organisers and mentors of the hackathon
A group from the Australian Structural Biology Computing (ASBC) community brought interdisciplinary expertise from UNSW’s Structural Biology Facility, WEHI, and the Monash AI Protein Design Program to optimise protein structure prediction and design workflows. The project incorporated GPU-enabled MMSeqs2 for accelerated sequence search and improved throughput and memory use of the Boltz-2 implementation for structure prediction. This work has been contributed to community-maintained Nextflow nf-core pipelines to promote efficient workflows for researchers worldwide.
A joint Australian Centre for Artificial Intelligence in Medical Innovation (La Trobe University) and Florey Institute of Neuroscience and Mental Health team tackled a highly complex 20,000-line codebase dedicated to childhood dementia research. Their project focused on predicting whether mRNA sequence data could accurately target the specific NPC1 mutation responsible for the disease. BioCommons’ AI Technical Lead, Dr Benjamin Goudey supported this work as a dedicated mentor. By bridging the gap between the hardware experts and the life science researchers, Ben helped the team navigate their massive codebase to ensure their mRNA dementia research workflows could run efficiently on the accelerated GPUs.
What technical outcomes were achieved?
By exploring batched inference strategies, some workloads in the structural biology project saw a four-fold speed increase, alleviating major predictive bottlenecks in high value protein design workflows. The team also doubled their maximum prediction size by chunking operations at key memory bottlenecks and working around overflow bugs in high-level libraries like PyTorch and trifast. With access to impressive hardware in the form of eight NVIDIA B200 GPUs, participating teams achieved substantial performance gains as well as insights into maximising the benefit of GPU acceleration within their research.
The event was a major upskilling opportunity, with researchers experimenting with agentic coding, getting introduced to GPU-accelerated data processing libraries (NVIDIA RAPIDS), and working under the guidance of expert mentors to iterate their optimisations.
Reflecting on the event, Dr Thomas Litfin, a UNSW researcher supported by Australian BioCommons and member of the ASBC community, highlighted the immense value of this hands-on support, as well as the lasting impact of the connections forged during the week:
‘The groups got an incredible amount of value from the expert mentors, who were instrumental in suggesting strategies and troubleshooting bottlenecks on the fly. The event was also a great community building exercise for our project by bringing together people from Monash, WEHI, and UNSW, and creating opportunities for ongoing collaborations.’
As AI methodologies become increasingly embedded in the life sciences, BioCommons will continue working alongside our partners to ensure researchers have both the cutting-edge digital infrastructure and the specialised skills needed to use it.
Read more about the event in the blog post by host Monash University.
Hear what the National AI Survey told BioCommons about researchers’ AI needs in the Future of AI in Life Sciences webinar.
Contributing to open and accessible training with 500+ bioinformatics resources
The Galaxy Training Network offers tutorials covering everything from genomics to machine learning. These self-paced resources combine videos and worked examples, allowing researchers to master complex bioinformatics tools.
Did you know that an international community has been coordinating the creation of freely accessible educational data analysis training resources for over a decade? This sustained stewardship from the Galaxy community has resulted in a collection of over 500 tutorials covering 35 topics that have been written, reviewed and maintained by people committed to open and accessible training.
What do these training resources cover?
Galaxy Training is a collection of tutorials developed and maintained by the worldwide Galaxy community. There have been 511 contributors, including the Hall of Fame: Australian BioCommons team who have contributed to 37 tutorials in support of life sciences computation. The library of tutorials available in Galaxy Training cover a wide range of scientific fields including genomics, climate science, ecology, statistics, machine learning and more. The resources combine videos, slides and worked examples that are accessible to everyone to use in their own time. The Galaxy community also hosts a massive online training event online each year to provide a further layer of support to learners.
Different categories of training resources available
The group of BioCommons contributors work at Melbourne Bioinformatics, the University of Queensland, and QCIF Digital Research, supporting researchers through training and improvements to the Galaxy data analysis platform. The Galaxy Australia service is made available for Australian life science researchers to more easily perform their data analysis. The web-accessible platform offers fully-subsidised accessible, reproducible, and transparent computational biological research. The Galaxy Australia service contains over 2,000 bioinformatics tools (for genome assembly, annotation, epigenetics, metabolomics, metagenomics, proteomics, statistics, transcriptomics, variant analysis and visualisation) and over 220 reference datasets (e.g. publicly available genome builds).
There are now over 650,000 registered Galaxy users internationally, and the Galaxy Community is always encouraging new users to get more involved. There are lots of different ways to contribute to the Galaxy Training Network (GTN), from reporting suggesting tune-up opportunities as you step through the tutorials, all the way to creating new content you think would be useful to others. Contributions can be driven by an individual sharing a specific thing they developed in the hope that it can help someone else with a similar niche interest, or involve a large effort to coordinate big changes that will benefit a very large group of people. The integration of the Galaxy Training Network with WorkflowHub was made possible thanks to a collaborative effort between Australian BioCommons and the Galaxy Training Network and WorkflowHub teams. All Galaxy Training workflows are now registered with WorkflowHub, with workflows contained in every new tutorial automatically pushed to WorkflowHub.
Where can I access Galaxy Training resources?
Investigate the Galaxy Training workflows: Galaxy Training on WorkflowHub.
View a webinar introduction to using Galaxy Australia for different biological applications: No code, no problem: data analysis for biologists using Galaxy Australia - Dr Tiff Nelson and Dr Tristan Reynolds.
Register for the free and online Galaxy Training Academy 2026.
Galaxy Australia’s 14 millionth job was a win for sustainable crop production
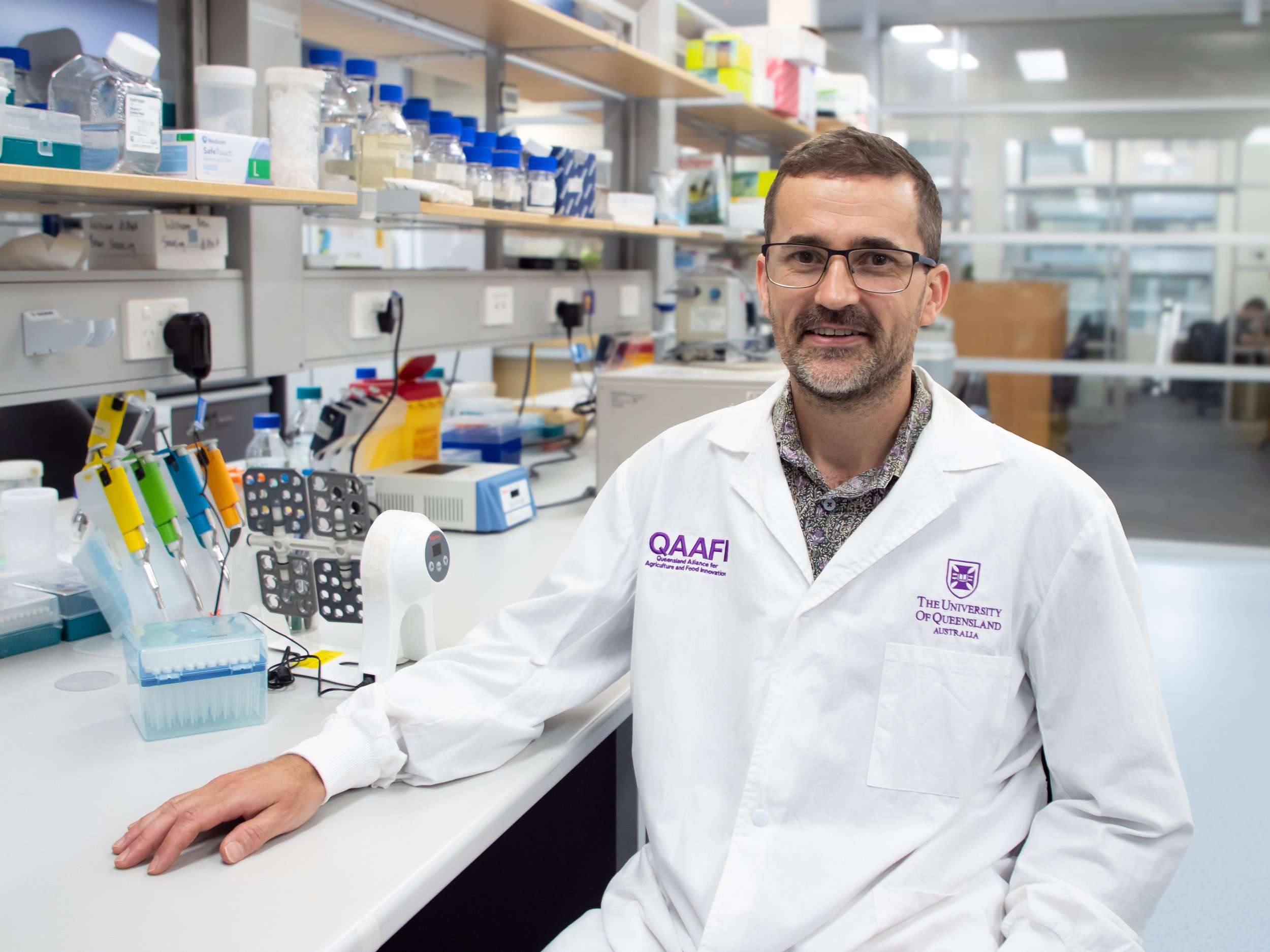
Galaxy Australia solves the technical bottleneck for molecular biologists who need to perform high-level bioinformatics without dedicated coding expertise. Molecular Biologist, Dr Donald Gardiner, from University of Queensland, uses the platform to understand how pathogens cause disease in plants.
The 14 millionth data analysis job was recently submitted to the Galaxy Australia platform, marking another major milestone for this national service. University of Queensland Senior Research Fellow, Dr Donald Gardiner, uses the platform to understand how pathogens cause disease in plants. His work highlights how essential access to the right research infrastructure is in sustaining Australia’s agricultural and biodiverse future.
How does Galaxy Australia support sustainable agriculture?
As part of the Queensland Alliance for Agriculture and Food Innovation, Donald works in a diverse group that includes molecular biologists, plant pathologists and biotechnologists to solve plant disease problems. Their work has been central to the ARC Hub for Sustainable Crop Protection, looking at innovative ways to protect Australian agriculture.
Dr Donald Gardiner
Donald’s work using Galaxy Australia spans several research projects, focusing on pathogens threatening the nursery and garden, forestry, agriculture and horticulture industries, as well as natural ecosystems. He is investigating the impact of Myrtle rust on the Mytaceae family of plants, including eucalypts and tea tree. The threat that pathogen Fusarium oxysporum poses to Australia’s banana industry, and risks to ginger and cereal crops are all better understood by Donald’s use of Galaxy Australia’s tools for genome assembly, annotation, RNA-seq and phylogenomics analysis.
How do molecular biologists use Galaxy Australia for bioinformatics?
For researchers like Donald whose lab work is complemented by key steps involving bioinformatics, the platform serves as a ‘truly enabling’ resource.
‘Galaxy allows me to skip much of the backend set up work for getting pieces of software to run. As someone who is not solely undertaking bioinformatic work, the need to have relatively simple ways to run programs is really important as I can’t dedicate massive amounts of time to bioinformatics.’
The service provides a comprehensive suite of software tools that are ready to use, allowing life science researchers to focus on the biological impact of their work rather than how to perform the analysis themselves, or relying on external bioinformatics expertise.
Find out more about Dr Donald Gardiner’s research
Learn about the different types of research that Galaxy Australia gets used for: No code, no problem - data analysis for biologists with Galaxy Australia (webinar)
Streamlined Science: Single sign-on with BioCommons Access
BioCommons Access launches in late March 2026 to provide a single sign-on for multiple analysis and data services across our ecosystem. This new system will streamline your research workflow by allowing seamless data transfers and unified access to platforms like Galaxy Australia, the Bioplatforms Australia Data Portal and more.
In a significant step towards integrating the tools and data that Australian life science researchers need, a new approach to accessing services across the Australian BioCommons ecosystem is coming. Launching in late March 2026, BioCommons Access will simplify access to multiple analysis and data services using a single sign-on. The availability of curated bundles of tools and data, and connections between services will streamline the research workflow.
BioCommons’ priorities are directed by the needs of life scientists, so services are always shaped by their research-driven stories. Rather than chasing new features down exciting tech rabbitholes, we nurture communities to hear researchers’ insights into how new services or better access to existing research infrastructures could transform their work.
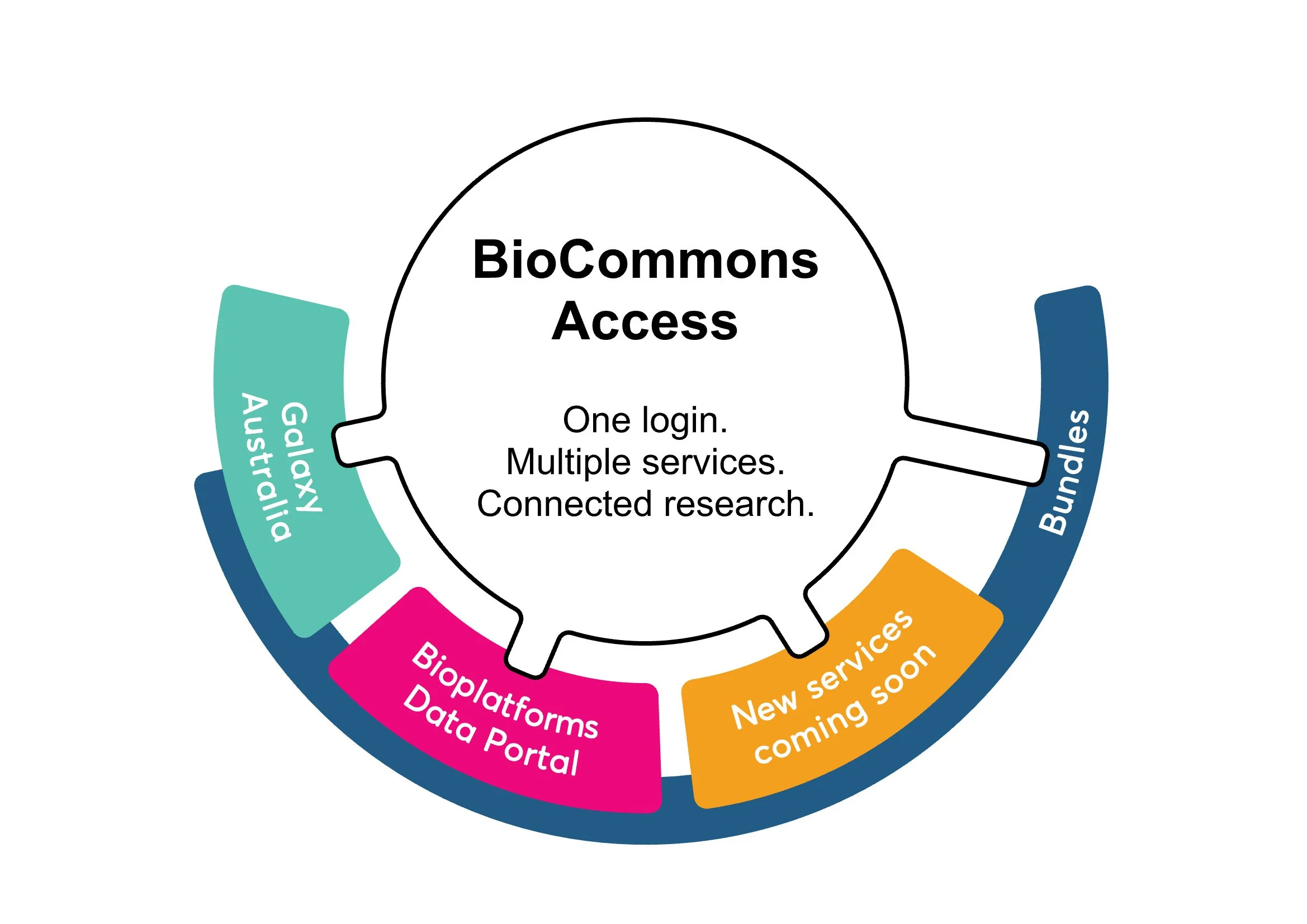
We’ve heard that researchers feel like they are forced to become the ‘network switch’ between the many isolated services they need to run increasingly complex workflows. Rather than managing multiple accounts, and manually jumping between different platforms, our BioCommons BioCloud team is building a linked ecosystem with a single sign-on that allows access to and transition between analysis and data services, including simple data transfer and increased visibility of available resources. Galaxy Australia and the Bioplatforms Australia Data Portal will be the first two services connected and available through BioCommons Access, with additional services to follow.
Australian BioCommons Access will enable:
Single sign-on access to multiple analysis and data services
Streamlined use of Galaxy Australia and the Bioplatforms Australia Data Portal using one login
A central portal to manage your user profile across services
Connection to curated bundles of tools and data
Fast one-click transfer of data between services (coming in May 2026).
The data and analysis needs of the Threatened Species Initiative (TSI) shaped the first service “Bundle” that is available as an add-on to BioCommons Access. When new members join this national consortium, they can sign up just once to start analysing the TSI data in the Bioplatforms Data Portal using Galaxy Australia. By adding the TSI Bundle to their registration, researchers are approved to access both embargoed and sensitive data as well as specialised and directly relevant analytics resources that are curated and managed by the TSI and Galaxy Australia.
BioCommons Access will be available from March 26th 2026, replacing the requirement for individual logins to Galaxy Australia and the Bioplatforms Data Portal. Existing users will be supported to migrate to the new system.
Future developments in BioCommons Access include the addition of institutional logins via the Australian Access Federation (AAF), new bundles to support national research communities, and additional integrated services.
The BioCommons Access service is the result of a deep collaboration between Galaxy Australia, the Bioplatforms Australia Data Portal, and the BioCloud team. BioCloud is the BioCommons division responsible for designing and operating the foundational and integration platforms that power molecular life sciences research in Australia.
Bringing together software engineers, cloud infrastructure specialists, and user experience experts based at the University of Melbourne and the University of Sydney, the BioCloud team builds secure, scalable platforms that enable researchers to work seamlessly across national capabilities. Expertise in digital identity and access management to deliver BioCommons Access was strengthened through a strategic partnership with BizData.
Help shape the future of AI in life sciences research and training
We are inviting community input to help shape the future of AI in life sciences. By participating in our survey and focus groups, you’ll help guide infrastructure priorities and shape our training program to benefit the Australian research community.
Artificial intelligence (AI) and machine learning (ML) are rapidly changing the way we work in life sciences. To ensure our community is well supported, Australian BioCommons is gathering input to map exactly how these technologies are being applied, particularly regarding molecular data analysis.
By sharing the current tools, processes, bottlenecks and skill gaps you experience, you will help prioritise national investments in digital infrastructure and directly shape our upcoming training programs.
How can you help shape the 2026 priorities?
We are looking for researchers and support staff at all levels of experience, from those who have never used AI to those who rely on it every day. Your perspective is valued regardless of your technical background.
There are two ways you can contribute:
Add your voice to the survey: Contribute to our national data collection to help identify trends and gaps across the Australian research ecosystem.
Join a focus group to contribute to our training programs: Share your specific challenges and ideas in a 60-minute interactive Zoom discussion. Register your interest by Friday 20 March 2026.
What happens next with your feedback?
We value your time and insights and we want to share what we learn along the way. In April, we will present a summary of our findings back with the community in a dedicated webinar. We will take a look at the initial training plan, giving you a sneak peek into the workshops and resources we’ll be launching later this year.
Accelerating deep learning in Australian structural biology research
The Australian Structural Biology Computing community has successfully translated the 2025 national research infrastructure roadmap into a year of impactful training, useful resources and global collaborations.
Enabled by rapid advances in deep learning methods for protein structure prediction, computational structural biology is driving major innovations in life sciences research. However, making effective use of these technologies requires cutting-edge software, highly specialised hardware, and interdisciplinary expertise.
To tackle these challenges, a passionate group of researchers officially formed the Australian Structural Biology Computing (ASBC) community in early 2025. Supported by BioCommons, this community-led initiative set out to share computational knowledge, methods and resources. Less than a year later, we can already see the value they bring to the Australian research sector.
How has the ASBC community grown?
Since its formation, the community has been moving quickly to address the needs identified in the Australian Structural Biology Deep-Learning Infrastructure Roadmap. The roadmap has already demonstrated its value to the sector, having been viewed over 1,100 times and downloaded over 900 times, in addition to being used in funding applications and publications.
To further connect researchers, BioCommons assisted in the development of a new Community Platform website that is now live, serving as a virtual hub for all users in Australia, with new resources and opportunities for contributions being added to the platform over time.
What training and resources are now available?
Capacity building has been a key focus for the ASBC community, and they have been integral to several key activities in 2025:
The community was instrumental in providing expertise for the ‘Leveraging deep learning to design custom protein-binding proteins’ webinar series, coordinated by BioCommons and featuring five individual guest speakers across the program of events
The facilitation of a 2025 ABACBS workshop ‘Leveraging peak Australian compute to enable workflows for predictive structural biology at scale’ to help researchers run protein prediction workflows on Australia’s peak high-performance computing (HPC) systems.
Collaborating with EMBL-EBI to develop the self-paced ‘Foundations in Structural Biology’ module (available now) for a global audience.
Get involved
Learn more and join the ASBC community
Investigate ways to engage with the community and its resources: https://www.biocommons.org.au/structural-biology
Roundtable convenes new community to chart Australia’s research infrastructure roadmap for AI in life sciences
Australian BioCommons and Bioplatforms Australia recently convened a national roundtable to establish a community of AI experts and researchers. This initiative leverages international partnerships with EMBL to build a unified strategic voice for Australia’s digital research infrastructure.
A roundtable to discuss the opportunities and implications of the use of AI in the life sciences was recently convened by Australian BioCommons and Bioplatforms Australia. It brought together almost 30 experts from diverse groups across Australia to share insights, experiences, and perspectives on AI in biosciences.
While helping to inform the strategic direction of Bioplatforms and BioCommons in this rapidly evolving area, the event also provided a valuable chance for researchers to hear from the Interim Director of EMBL European Bioinformatics Institute, Dr Johanna McEntyre, about how Europe is approaching our shared challenges. Given the Memorandum of Understanding between Australian BioCommons, Bioplatforms Australia and the European Molecular Biology Laboratory (EMBL), it was a wonderful opportunity to reflect on the value of international collaboration in this space while Jo was visiting Australia.
(L-R) Jeff Christiansen, Johanna McEntyre and Benjamin Goudey
While research infrastructure was the focus, a varied group of people who use, develop or support AI in Australian life science research participated in presentations, panel discussions and interactive exercises. People from universities, research institutes, computation service providers and other NCRIS facilities shared how they are using and building AI, and learned about needs, gaps, and constraints to their work.
The roundtable was a step towards finding strategic partners who can help to map the current landscape, identify gaps and define investment priorities. As our AI in Molecular Life Sciences community coalesces, we can expect to gain a deep understanding of the challenges facing Australian life scientists using and developing AI tools and models, and benefit from having a diverse partner network acting as a collective watchtower for trends and opportunities that can define the new shape of life sciences computing.
If you have insights into how AI will impact research methods and future discoveries, please consider contributing to the development of a BioCommons Infrastructure Roadmap for AI in the Life Sciences by engaging with this newly formed group.
Sign up to join a new community of people who will work together to identify and document the infrastructure challenges and investment priorities for Australian life sciences
Other opportunities to help shape the future of AI in life sciences research and training in Australia