News
Subscribe to the Australian BioCommons monthly newsletter or read previous editions
Contributing to open and accessible training with 500+ bioinformatics resources
The Galaxy Training Network offers tutorials covering everything from genomics to machine learning. These self-paced resources combine videos and worked examples, allowing researchers to master complex bioinformatics tools.
Did you know that an international community has been coordinating the creation of freely accessible educational data analysis training resources for over a decade? This sustained stewardship from the Galaxy community has resulted in a collection of over 500 tutorials covering 35 topics that have been written, reviewed and maintained by people committed to open and accessible training.
What do these training resources cover?
Galaxy Training is a collection of tutorials developed and maintained by the worldwide Galaxy community. There have been 511 contributors, including the Hall of Fame: Australian BioCommons team who have contributed to 37 tutorials in support of life sciences computation. The library of tutorials available in Galaxy Training cover a wide range of scientific fields including genomics, climate science, ecology, statistics, machine learning and more. The resources combine videos, slides and worked examples that are accessible to everyone to use in their own time. The Galaxy community also hosts a massive online training event online each year to provide a further layer of support to learners.
Different categories of training resources available
The group of BioCommons contributors work at Melbourne Bioinformatics, the University of Queensland, and QCIF Digital Research, supporting researchers through training and improvements to the Galaxy data analysis platform. The Galaxy Australia service is made available for Australian life science researchers to more easily perform their data analysis. The web-accessible platform offers fully-subsidised accessible, reproducible, and transparent computational biological research. The Galaxy Australia service contains over 2,000 bioinformatics tools (for genome assembly, annotation, epigenetics, metabolomics, metagenomics, proteomics, statistics, transcriptomics, variant analysis and visualisation) and over 220 reference datasets (e.g. publicly available genome builds).
There are now over 650,000 registered Galaxy users internationally, and the Galaxy Community is always encouraging new users to get more involved. There are lots of different ways to contribute to the Galaxy Training Network (GTN), from reporting suggesting tune-up opportunities as you step through the tutorials, all the way to creating new content you think would be useful to others. Contributions can be driven by an individual sharing a specific thing they developed in the hope that it can help someone else with a similar niche interest, or involve a large effort to coordinate big changes that will benefit a very large group of people. The integration of the Galaxy Training Network with WorkflowHub was made possible thanks to a collaborative effort between Australian BioCommons and the Galaxy Training Network and WorkflowHub teams. All Galaxy Training workflows are now registered with WorkflowHub, with workflows contained in every new tutorial automatically pushed to WorkflowHub.
Where can I access Galaxy Training resources?
Investigate the Galaxy Training workflows: Galaxy Training on WorkflowHub.
View a webinar introduction to using Galaxy Australia for different biological applications: No code, no problem: data analysis for biologists using Galaxy Australia - Dr Tiff Nelson and Dr Tristan Reynolds.
Register for the free and online Galaxy Training Academy 2026.
Galaxy Australia’s 14 millionth job was a win for sustainable crop production
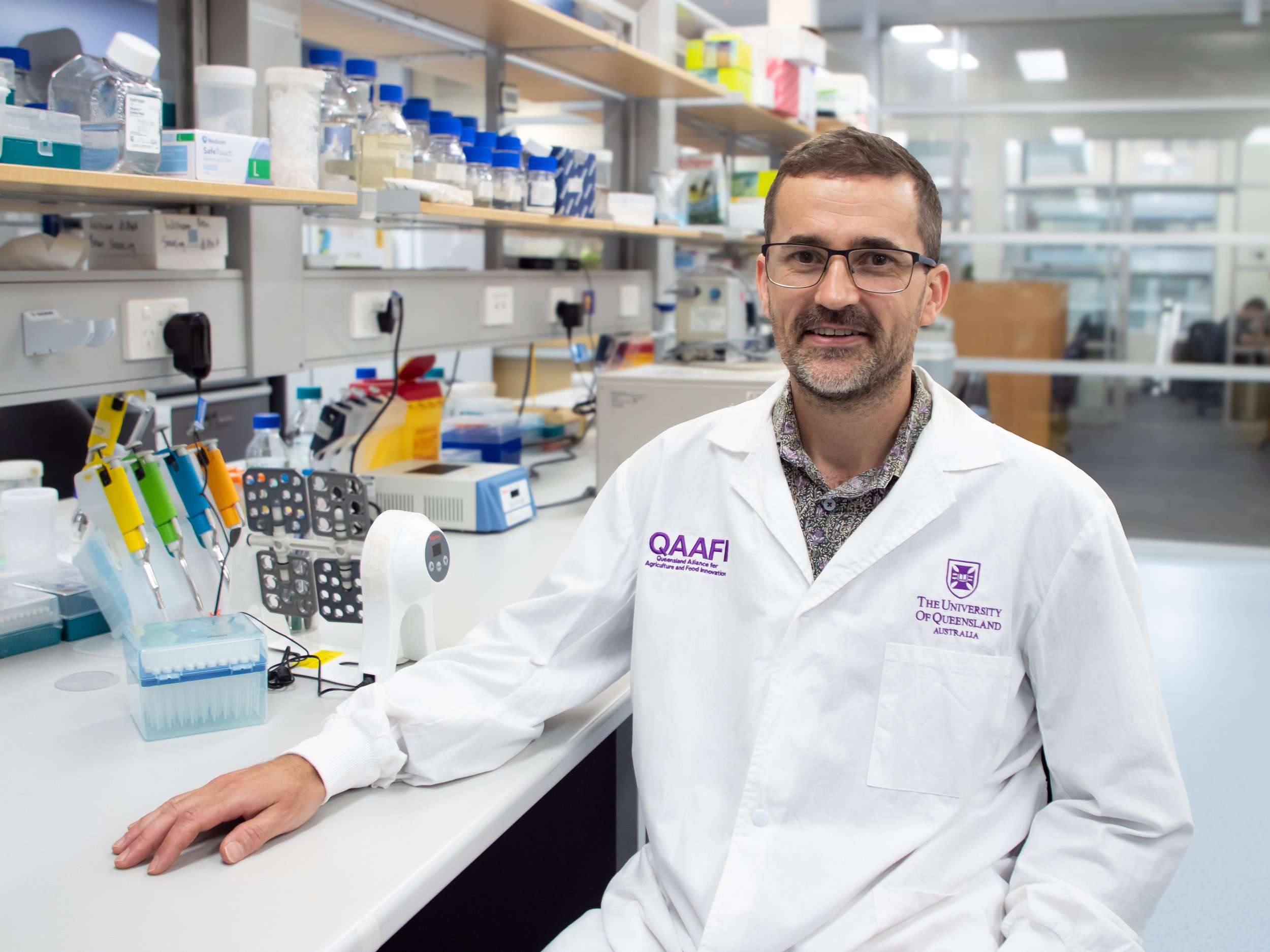
Galaxy Australia solves the technical bottleneck for molecular biologists who need to perform high-level bioinformatics without dedicated coding expertise. Molecular Biologist, Dr Donald Gardiner, from University of Queensland, uses the platform to understand how pathogens cause disease in plants.
The 14 millionth data analysis job was recently submitted to the Galaxy Australia platform, marking another major milestone for this national service. University of Queensland Senior Research Fellow, Dr Donald Gardiner, uses the platform to understand how pathogens cause disease in plants. His work highlights how essential access to the right research infrastructure is in sustaining Australia’s agricultural and biodiverse future.
How does Galaxy Australia support sustainable agriculture?
As part of the Queensland Alliance for Agriculture and Food Innovation, Donald works in a diverse group that includes molecular biologists, plant pathologists and biotechnologists to solve plant disease problems. Their work has been central to the ARC Hub for Sustainable Crop Protection, looking at innovative ways to protect Australian agriculture.
Dr Donald Gardiner
Donald’s work using Galaxy Australia spans several research projects, focusing on pathogens threatening the nursery and garden, forestry, agriculture and horticulture industries, as well as natural ecosystems. He is investigating the impact of Myrtle rust on the Mytaceae family of plants, including eucalypts and tea tree. The threat that pathogen Fusarium oxysporum poses to Australia’s banana industry, and risks to ginger and cereal crops are all better understood by Donald’s use of Galaxy Australia’s tools for genome assembly, annotation, RNA-seq and phylogenomics analysis.
How do molecular biologists use Galaxy Australia for bioinformatics?
For researchers like Donald whose lab work is complemented by key steps involving bioinformatics, the platform serves as a ‘truly enabling’ resource.
‘Galaxy allows me to skip much of the backend set up work for getting pieces of software to run. As someone who is not solely undertaking bioinformatic work, the need to have relatively simple ways to run programs is really important as I can’t dedicate massive amounts of time to bioinformatics.’
The service provides a comprehensive suite of software tools that are ready to use, allowing life science researchers to focus on the biological impact of their work rather than how to perform the analysis themselves, or relying on external bioinformatics expertise.
Find out more about Dr Donald Gardiner’s research
Learn about the different types of research that Galaxy Australia gets used for: No code, no problem - data analysis for biologists with Galaxy Australia (webinar)
Streamlined Science: Single sign-on with BioCommons Access
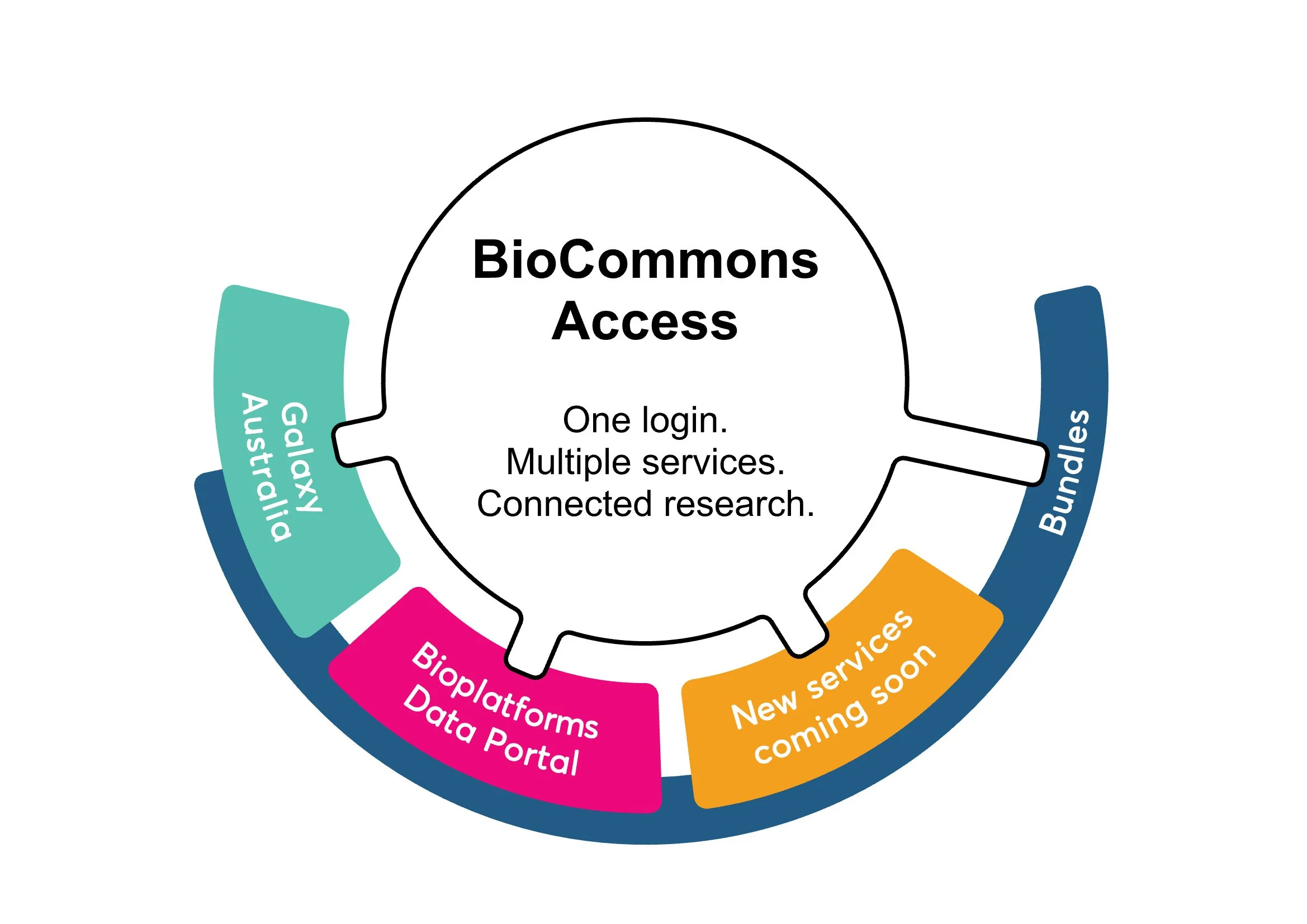
BioCommons Access launches in late March 2026 to provide a single sign-on for multiple analysis and data services across our ecosystem. This new system will streamline your research workflow by allowing seamless data transfers and unified access to platforms like Galaxy Australia, the Bioplatforms Australia Data Portal and more.
In a significant step towards integrating the tools and data that Australian life science researchers need, a new approach to accessing services across the Australian BioCommons ecosystem is coming. Launching in late March 2026, BioCommons Access will simplify access to multiple analysis and data services using a single sign-on. The availability of curated bundles of tools and data, and connections between services will streamline the research workflow.
BioCommons’ priorities are directed by the needs of life scientists, so services are always shaped by their research-driven stories. Rather than chasing new features down exciting tech rabbitholes, we nurture communities to hear researchers’ insights into how new services or better access to existing research infrastructures could transform their work.
We’ve heard that researchers feel like they are forced to become the ‘network switch’ between the many isolated services they need to run increasingly complex workflows. Rather than managing multiple accounts, and manually jumping between different platforms, our BioCommons BioCloud team is building a linked ecosystem with a single sign-on that allows access to and transition between analysis and data services, including simple data transfer and increased visibility of available resources. Galaxy Australia and the Bioplatforms Australia Data Portal will be the first two services connected and available through BioCommons Access, with additional services to follow.
Australian BioCommons Access will enable:
Single sign-on access to multiple analysis and data services
Streamlined use of Galaxy Australia and the Bioplatforms Australia Data Portal using one login
A central portal to manage your user profile across services
Connection to curated bundles of tools and data
Fast one-click transfer of data between services (coming in May 2026).
The data and analysis needs of the Threatened Species Initiative (TSI) shaped the first service “Bundle” that is available as an add-on to BioCommons Access. When new members join this national consortium, they can sign up just once to start analysing the TSI data in the Bioplatforms Data Portal using Galaxy Australia. By adding the TSI Bundle to their registration, researchers are approved to access both embargoed and sensitive data as well as specialised and directly relevant analytics resources that are curated and managed by the TSI and Galaxy Australia.
BioCommons Access will be available from March 26th 2026, replacing the requirement for individual logins to Galaxy Australia and the Bioplatforms Data Portal. Existing users will be supported to migrate to the new system.
Future developments in BioCommons Access include the addition of institutional logins via the Australian Access Federation (AAF), new bundles to support national research communities, and additional integrated services.
The BioCommons Access service is the result of a deep collaboration between Galaxy Australia, the Bioplatforms Australia Data Portal, and the BioCloud team. BioCloud is the BioCommons division responsible for designing and operating the foundational and integration platforms that power molecular life sciences research in Australia.
Bringing together software engineers, cloud infrastructure specialists, and user experience experts based at the University of Melbourne and the University of Sydney, the BioCloud team builds secure, scalable platforms that enable researchers to work seamlessly across national capabilities. Expertise in digital identity and access management to deliver BioCommons Access was strengthened through a strategic partnership with BizData.
Making Microbiome workflows more accessible for researchers
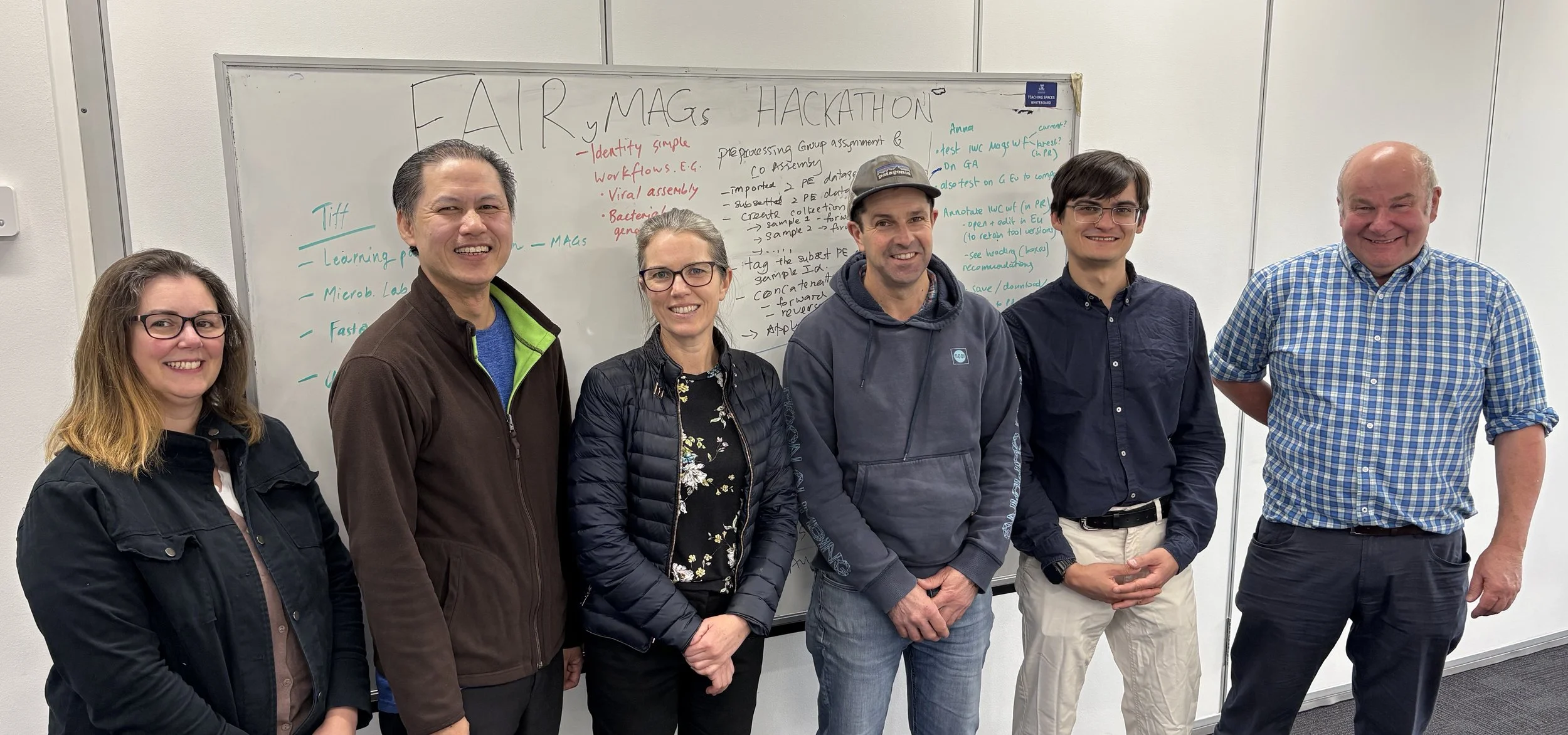
A group of Australian microbiome specialists recently joined multiple ELIXIR nodes and the international Galaxy community for an ‘Optimising MAGS-building workflows’ hackathon.
A group of Australian microbiome specialists recently participated in the Optimising MAGs-building workflows hackathon, joining multiple ELIXIR nodes and the international Galaxy community in Europe. Gathering to work together from Melbourne and connecting online with international colleagues, the group worked on enhancing the FAIRness of MAGs-building workflows and developing user-friendly training materials to support their uptake.
Microbiology, microbiome, and metagenomics researchers generate Metagenome Assembled Genomes (MAGs) from samples of mixed DNA in soil, air, water or other sources. Sophisticated workflows for genome assembly, including the scoring of completeness and contamination are required for identification of what is present in the sample and an indication of its function.
The Hackathon Team (L-R): Anna Syme, Mike Thang, Tiff Nelson, Andrew Bisset, Viní Salazar, Dieter Baluch
There are a few trusted workflows that researchers rely on like nf-core/mag and the Galaxy Metagenome-Assembled Genomes (MAGs) Generation workflow. This event focussed on improving Galaxy MAGs workflows and developing self-guided materials to add to the Galaxy Training collection of tutorials.
Assoc Prof Dieter Bulach joined the event to better understand if he might be able to use this MAGs workflow on his project’s data. As a Bioinformation in the Ecology & Environment Team of the Revitalising Informal Settlements and their Environments (RISE), he has metagenomic data with lots of potential. Based in the Melbourne Veterinary School, Dieter is looking at antimicrobial resistance in bacteria, to support the RISE consortium’s vision to transform human and environmental health through water-sensitive changes.“I run training for wet lab researchers who are actively learning bioinformatics while they are doing assemblies. Improving the Galaxy MAGs workflow and training resources will help them use these tools. And being at the hackathon gives me connections into the Galaxy project, exposure to new capabilities in the platform, visibility of what’s going on in Europe and importantly, who to ask when I need help!”
Direct assistance was available from two Galaxy Australia team members who were participating: Dr Anna Syme (Australian BioCommons) and Michael Thang (University of Queensland). Given both are passionate about supporting researchers and regularly offer training in bioinformatics techniques, their insights were invaluable to the hackathon group. Melbourne Bioinformatics’ Vini Salazar also brought an education focus, given he is a subject coordinator in the University of Melbourne MSci (Bioinformatics) program. With extensive experience in metagenomics and software education, Vini worked on updating existing training materials to reflect current best practices in MAG-building.
Dr Andrew Bissett participated with a different goal in mind. As the group leader of the Marine Observations and Analytics in CSIRO's Sustainable Marine Futures program, and the Scientific Lead of the Australian Microbiome Initiative, he was interested in raising awareness of valuable microbial genomic data. By referencing the Australian Microbiome data in Galaxy training materials, Andrew hopes to highlight the availability of this unique resource to a new international audience.
While everyone brings different skills and motivations to our hackathons, the overwhelming feedback is that the time spent together in person leads to wide-ranging conversations, new connections and unexpected benefits. Community Engagement Lead for BioCommons, Dr Tiff Nelson, identified the potential for this hackathon to bring together people who otherwise wouldn’t collaborate on shared tasks, and to drive international activities with local benefits. It is just one activity of many that supports the research community’s vision to help researchers undertake microbiome analyses that was captured in the Microbiome Analysis Infrastructure Roadmap for Australia.
Thousands join international Galaxy training events
The Galaxy platform is celebrating its 20th anniversary this year and Galaxy Australia is one of the core BioCommons services. We are proud to provide and support a variety of opportunities for researchers to engage with the team, community and tools.
The Galaxy platform is celebrating its 20th anniversary this year. This collaborative data analysis platform is widely used by scientists around the world, and underpins many computational biology services. Galaxy Australia is one of the core services BioCommons delivers, so naturally we provide lots of opportunities for researchers to engage with the team, community and tools.
The Galaxy Australia team recently supported the Singapore Biology League, who chose to incorporate Galaxy into their online collaborative biology contest for the first time. This massive event welcomed over 2000 pre-university students who formed teams to run 2054 tools on Galaxy Australia over 4 hours. It was a great opportunity to broaden participants’ exposure to biology and bioinformatics beyond the school curriculum.
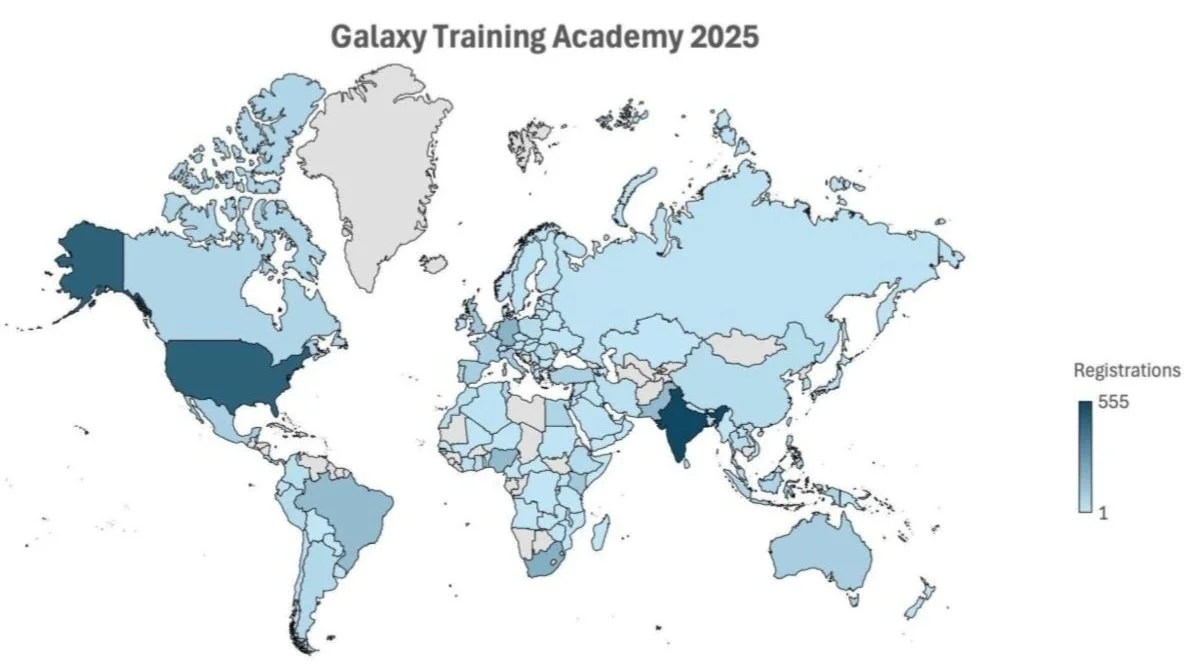
The week-long Galaxy Training Academy attracted more than 3500 people from across the globe this year. Seventy three Australian researchers were amongst the international cohort joining from their homes and offices to work through tutorials, with support as they needed it. Live help was provided in our time zones by Galaxy Australia’s Dr Anna Syme, Dr Igor Makunin and Dr Tristan Reynolds, who fielded questions on a wide range of topics from fungal genomics to bat ecology! Participants brought their own specialities as they learned how to use the fully subsidised Galaxy Australia platform for proteomics, genome assembly, transcriptomics, single cell RNAseq, microbiome analysis, machine learning and more.
In the lead up to the Galaxy Training Academy, BioCommons’ Dr Tiffanie Nelson and Galaxy Australia’s Dr Tristan Reynolds helped researchers understand what’s possible by presenting the webinar: No code, no problem: data analysis for biologists with Galaxy Australia. Examples included how Galaxy Australia is being used for biosecurity screening, foodborne pathogen detection and building reference genomes of the critically endangered swift parrot, as well as a tour of the practical features of Galaxy Australia that make sophisticated workflows like these accessible to all, regardless of their computational skills.
Get started by exploring the tutorials available: Galaxy Training Network
If you’re an existing Galaxy Australia user, improve your methods with our Top Tips videos
Stop the rot! Plant Bacteriologist’s genome assembly becomes Galaxy Australia’s 11 millionth job
An enthusiastic new user recently submitted the lucky 11 millionth data analysis job to the Galaxy Australia platform. Plant Bacteriologist Dr Toni Chapman has begun regularly using the fully-subsidised service for her genome assemblies of bacteria important to agricultural plant biosecurity and production.
Image courtesy of Dr Toni Chapman
An enthusiastic new user recently submitted the lucky 11 millionth data analysis job to the Galaxy Australia platform. Plant Bacteriologist Dr Toni Chapman has begun regularly using the service for her genome assemblies of bacteria important to agricultural plant biosecurity and production.
As a Senior Research Scientist in Agriculture and Biosecurity at the NSW Department of Primary Industries and Regional Development (DPIRD), Toni’s work spans both diagnostics and research. Using NovaSeq and HiFi sequencing technologies, she assembles bacterial genomes to help growers diagnose the cause of disease symptoms in their plants or to identify bacteria present in suspect samples collected by biosecurity officers.
In the past, Toni needed to rely heavily on bioinformaticians for assistance with genome assembly of the bacteria she works with. Now after attending a training workshop and signing up for access to the fully-subsidised Galaxy Australia service, she independently completes more of the analysis steps and is able to identify the bacterial pathogens herself.
“Over time I have been learning to code and to use various programs for genome assembly, but only returning to it on a casual basis makes it very hard to maintain the necessary skills in the bioinformatics space. Being able to design workflows in Galaxy Australia that I can come back to at any time makes assembling genomes easy, even if there has been weeks or months between visits.”
Toni contributes to the Plant Pathogen ‘Omics Initiative, which is generating molecular reference data for plant pathogens in Australia. Established by Bioplatforms Australia to support research in plant protection and enhance biosecurity surveillance efforts, the national plant pathogen community is collaborating to create high quality data that can be shared with all researchers via the Data Portal. Toni was one of a large group that came together with members of the Functional Fungi and Plant Pathogen ‘Omics National Initiatives for a hands-on bioinformatics workshop, learning how to use Galaxy and the programs needed for genome assembly using their own real-world scenarios.
As part of her contribution to the Initiative, Toni is conducting genome assembly on Pseudomonas species that cause disease in plants, to update the taxonomy of collection isolates and gain a more accurate view of which pathogens are in Australia. In another project, she is sequencing the bacterial pathogens that cause soft rot disease in plants. The incidence of soft rot is increasing, and so is the range of bacteria that can cause the disease. The newly assembled genomes of these bacteria are updating existing culture collection identities and helping to understand the diversity of bacteria that cause soft rot infections in Australia.
If you’d like to find out more about her work, see Dr Toni Chapman’s publications.
If you are interested in hearing about the different types of research that Galaxy Australia gets used for, watch this recorded webinar: No code, no problem - data analysis for biologists with Galaxy Australia.
An Australian community for computational structural biology
A passionate group of structural biologists has formed the Australian Structural Biology Computing Community, to share computational knowledge, methods, and resources.
The active new community is receiving support from a range of partners and advocates, including L-R Johan Gustafsson (BioCommons), Steven Manos (BioCommons), Kate Michie (UNSW) and Andrew Gilbert (Bioplatforms Australia)
The explosion of possibilities presented by deep learning approaches in structural biology research has created many new opportunities and challenges. A passionate group of structural biologists has formed the Australian Structural Biology Computing Community, to approach this new era as part of a community that shares computational knowledge, methods, and resources.
This community-driven approach brings together a diverse group of people, with initial contributions forming around leads from the Structural Biology Facility at UNSW, and an academic panel of experts from Monash University, Walter and Eliza Hall Institute of Medical Research (WEHI), University of Western Australia (UWA), Australian National University (ANU), Bio21 Institute of Molecular Science and Biotechnology (Bio21), University of Melbourne, La Trobe University, University of Queensland (UQ) - IMB, University of Sydney, Griffith University, Swinburne University of Technology, CSIRO, and the University of Adelaide. Anyone involved in structural biology in Australia is invited to join and there are lots of different ways to get involved.
The Community for Structural Biology Computing in Australia webpage is a useful new resource for all users of computing for structural biology research in Australia. The page is constantly evolving and expanding, and it currently focuses on the use of deep learning methods in Structural Biology. It includes practical guides on topics like “Best practices for presenting and sharing AlphaFold models in a paper” as well as news items and announcements for relevant courses and meetings.
Australian BioCommons supports the community by hosting quarterly online meetings that aim to tease out how computational structural biologists’ challenges might be addressed with community-scale responses and national research infrastructure solutions. If you join the mailing list via the community webpage, you will receive updates and invitations to community meetings and the discussions in Slack.
BioCommons began providing broad, fully subsidised, access to structural prediction in 2022 by making AlphaFold2 available within its Galaxy Australia service. The Australian AlphaFold2 Service provides both an easy-to-use interface and dedicated GPUs to Australian researchers. When BioCommons hosted the international 2023 Galaxy Community Conference, the keynote speech by Chief Scientist of the Structural Biology Factility at UNSW, Kate Michie, generated much excitement around forming an Australian community of practice for computational structural biology as an avenue for collectively addressing the challenges presented by deep learning in structural biology.
BioCommons has supported key research stakeholders to refine the new community’s purpose, began running quarterly community meetings, and helped to establish the shared community spaces like the Australian Structural Biology Computing website and GitHub. As well as facilitating consultations with infrastructure partners and the broader computational infrastructure community, a group of national panel of experts has been identified.
This community collaborates with their peers to:
Collectively create and maintain community forums and centralised collaboration platforms to support collaboration and knowledge sharing (i.e. methods and documentation);
Foster collaboration between structural biologists, computer scientists, and data scientists, thereby creating interdisciplinary teams to help tackle complex challenges, validate results and ensure robust applications of deep learning methods;
Lead the review, prioritisation, testing, optimisation, and sharing of deep learning codes, software and approaches that are of broad relevance and interest to the Australian research community;
Develop quality assessment tools to help evaluate the quality of calculated structures, and help guide researchers towards reliable predictions; and,
Address the ethical implications of AI-driven structural predictions, as well as discuss transparency, bias and interpretability to ensure responsible use of these technologies.
The Australian Structural Biology Community is poised to tackle a set of pilot activities aimed at fast tracking a national response to the challenges facing computational approaches in structural biology. A much anticipated future output is an infrastructure roadmap document that will formalise and describe the high level requirements of the community. This collaborative effort between the new Australian Structural Biology Community, Australian BioCommons, and BioCommons infrastructure partners will support the Australian Structural Biology community as new needs arise relating to bioinformatics tools, software, infrastructure or training.
Keep in touch by subscribing for updates at the Community for Structural Biology Computing in Australia webpage.
Repurposed hardware boosts national capacity and powers innovation
QCIF Ltd has made high-performance hardware available to the Australian BioCommons, giving the hardware a second life and uplifting national capacity for running AlphaFold 2 jobs in Galaxy Australia while supporting innovation through other GPU-enabled tools.
This story is co-published with QCIF Ltd
After successfully completing a previous project, QCIF Ltd made available high-performance hardware to the Australian BioCommons, giving the hardware a second life in enabling research and uplifting national capacity for the benefit of the scientific community.
Well suited for running AlphaFold 2 jobs, the five General-Purpose Graphics Processing Units (GPGPUs) are now being used to enhance the national compute network behind the Galaxy Australia service.
The impact of this repurposing goes beyond infrastructure improvements. It has significantly expanded Galaxy Australia's capacity to support research and innovation by enabling the use of other GPU-enabled tools that offer major benefits to the scientific community. GPU processing can provide massive improvements in computational efficiency, decreasing processing times to less than 5% of conventional equivalents.
Dr Cameron Hyde, a bioinformatician at QCIF who supports the development of national software platforms like Australian BioCommons' Galaxy and Apollo services, co-authored the original AlphaFold 2 wrapper that enabled the tool to run within Galaxy Australia, ensuring both a friendly user-interface as well as instant access to the GPU clusters required to power the tool. He shared his enthusiasm for the new possibilities unlocked by the repurposed hardware which was originally part of an investment made in 2021 by the Australian Research Data Commons (ARDC) to support national platform projects and now directly enhances the bioinformatics services he helps deliver to Australian researchers. “Now that we have five GPU nodes of our own, we have room to experiment and explore new GPU-enabled tools. This gives us room to innovate beyond AlphaFold and accelerate scientific discovery in other research domains.”
For example, Galaxy Australia’s lead Bioinformatician Michael Thang has been using the hardware to explore running Nanopore’s “Dorado” on Galaxy Australia. Dorado is a high-performance basecaller for Oxford Nanopore Technology sequencing data. This innovation would enable researchers to conduct their entire analysis, from raw sequencing data through to assembled genome, all within the Galaxy Australia service.
Collaboration driving innovation
Developed by Google DeepMind, AlphaFold is an AI system that predicts a protein’s 3D structure from its amino acid sequence with accuracy comparable to experimental methods. In 2020, Australian BioCommons identified an opportunity to democratise access to this powerful tool by making AlphaFold 2 available through Galaxy Australia. This gave Australian researchers much greater accessibility to AlphaFold 2, allowing life scientists to easily visualise proteins in a manner inaccessible to all but dedicated structural biology researchers. This advance has supported research into protein-protein interactions, activation and inhibition mechanisms, and drug design.
By 2025, use of AlphaFold 2 has surged, evolving from an analytical tool for individual proteins into a routine screening tool for studying protein-protein interactions. To support this shift, Dr Hyde collaborated closely with Australian Structural Biology Computing Community to develop extensions to the AlphaFold Galaxy tool, including new output formats, input parameters, and an option to re-use intermediate files for improved efficiency.
Supported by the Australian BioCommons, AARNnet, QCIF Ltd, and The University of Melbourne, the optimised system now provides fully subsidised access for all Australian researchers via the Australian Alphafold Service. We extend our sincere thanks to the Australian Research Data Commons (ARDC) for providing the hardware to QCIF Ltd and enabling its reuse by Australian BioCommons.
Microbiology Lab empowers microbial data analysis with integrated tools, workflows and compute
Microbiology Lab offers a customised, user-friendly view of Galaxy Australia that provides rapid access to popular tools, workflows and compute for analysis and visualisation of microbiomes and omics data from microbial isolates.
Galaxy Australia’s Microbiology Lab is now providing researchers with rapid access to popular tools, workflows and compute for analysis and visualisation of microbiomes and omics data from microbial isolates.
Development of this curated view of Galaxy Australia was driven by the international microbial research community who identified the most commonly used tools in the field through meticulous research, surveys and community consultations.
The Lab integrates more than 220 tools and 65 workflows with step-by-step tutorials and structured learning paths for a suite of different analyses. Microbiology Lab pairs perfectly with the computing power of Galaxy Australia, which is underpinned by computational resources provided by AARNet, ARDC Nectar Research Cloud, the University of Melbourne, QCIF, Pawsey Supercomputing Research Centre, National Computational Infrastructure, and Microsoft Azure. From raw data, to differential analysis, visualisation and assembly, Microbiology Lab makes it easy to get started with reproducible analysis of microbial data.
Galaxy Australia’s Microbiology Lab is the latest release in a series of Labs supporting research domains. It represents an important step forward in Australian BioCommons activities to support Australian microbial research. The Microbiome Analysis Infrastructure Roadmap for Australia identified a need to implement a shared platform that eases access to preferred tools and workflows for analysis of microbial data with sufficient computational power. This fully-subsidised resource is expected to improve efficiency for researchers.
If you are an Australian researcher with an interest in microbial isolates and microbiomes, be sure to take a tour of the Galaxy Australia Microbiology Lab and try it out now!
Join 41,000 users of this fully-subsidised and easy to use Australian bioinformatics platform
Galaxy has proven to be such a versatile data analysis platform that Galaxy Australia supports over 41,000 users. Now is the perfect time to get on board with Galaxy, with a variety of ways to get involved on offer in the coming months.
Galaxy has proven to be such a versatile data analysis platform that Galaxy Australia now supports over 41,000 users. This fully subsidised, open-source system for analysing and visualising data, authoring workflows, training, publishing tools, and managing infrastructure is powered by a world-wide community of 500,000 Galaxy users as well as the institutions who invest in its growth. Now is the perfect time to get on board with Galaxy, with a variety of ways to get involved on offer in the coming months.
If you are curious to see how Galaxy is being used in a wide range of research settings, come along to our upcoming webinar showcasing how Galaxy Australia is supporting genome assembly and annotation, metagenomics, proteomics, transcriptomics, data visualisation and more. Register now for No code, no problem - data analysis for biologists with Galaxy Australia on Tue 29 Apr 2025.
The global community’s annual training event - Galaxy Training Academy - is coming up in May. This week-long, completely free, online training event will help you master the Galaxy platform for data analysis. It is self-paced and you can choose from topics including Proteomics, Assembly, Transcriptomics, Single Cell, Microbiome or Machine Learning data analysis in Galaxy. The Galaxy Australia team will be on hand to answer any questions you have during the event over 12-16 May.
Most people simply make use of the platform to do their research, but there’s an open invitation to contribute to this lively community too. The international Galaxy Community,has put together a Get Started page to help you find out more about terminology, resources, servers, types of communities, and the different ways to contribute to the great things the Galaxy community achieves together. You can also connect with the community in person at the Galaxy and Bioconductor Community Conference 2025 is coming up in June in the USA.
Strategic partnerships formed around the platform are key in making it readily available to Australian researchers. Australian BioCommons is able to offer the service at no cost to researchers because of the commitment of valuable partners QCIF, The University of Melbourne and AARNet who help manage the Galaxy Australia service and provision of computational resources provided by respected national institutions and providers AARNet, ARDC Nectar Research Cloud, the University of Melbourne, QCIF, National Computational Infrastructure, Pawsey Supercomputing Research Centre and Microsoft Azure. These efforts are backed by funding from the University of Melbourne, the Queensland state government and Bioplatforms Australia, who bring Federal investment through the National Collaborative Research Infrastructure Strategy.
If you’re ready to start using Galaxy Australia, you can jump right in and try it now: usegalaxy.org.au/