News
Subscribe to the Australian BioCommons monthly newsletter or read previous editions
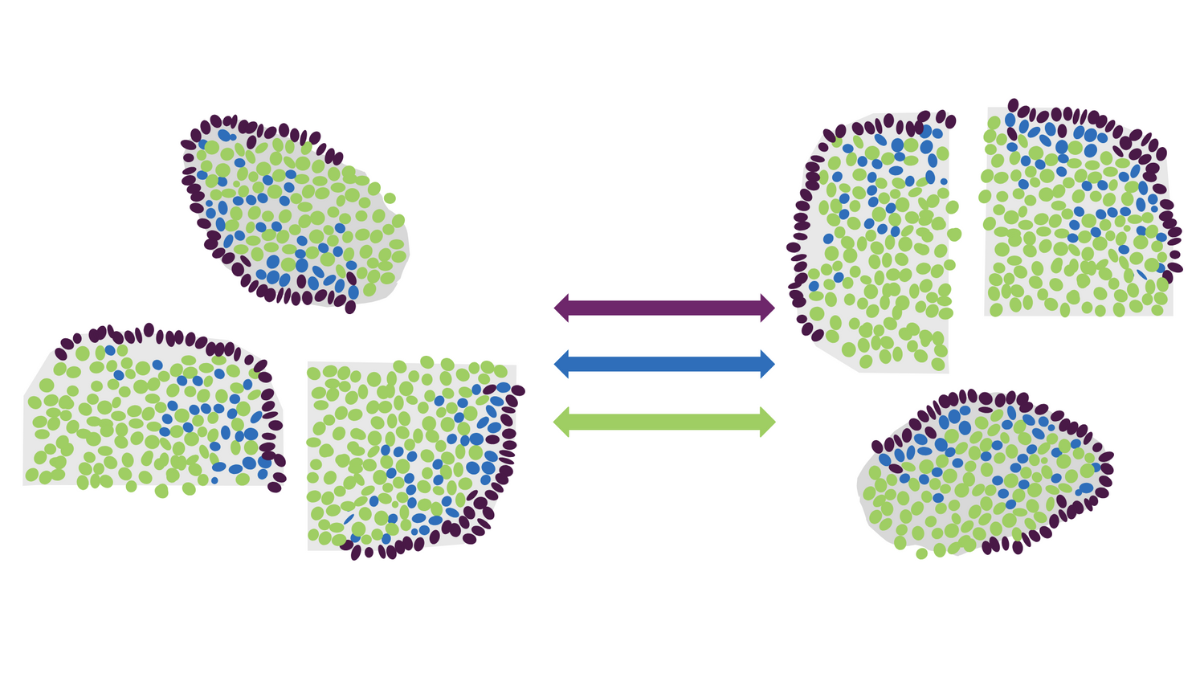
Introducing Spatial Sampler, the ‘cheat sheet’ to answering biological questions
A new tool is available to help researchers quickly answer important questions about their spatial omics data. This new resource, called Spatial Sampler, collates example R scripts for a variety of comparative analyses to fast track analysis.
A new tool is available to help researchers quickly answer important questions about their spatial omics data. This new resource, called Spatial Sampler, collates example R scripts for a variety of comparative analyses to fast track analysis.
QCIF Digital Research Senior Bioinformatician, Dr Sarah Williams, developed the resource for her own use, and upon realising its potential value to others she went on to share the collection of examples via the BioCommons’ Workflow Commons project.
What is Spatial Sampler?
Spatial Sampler is a collection of ‘How-To’ guides framed around testing specific biological questions.
Each How-To includes:
An example workflow using a real dataset, which explains why each step is necessary
What input data is needed
What the output looks like
A code snippet that can be copy-pasted
Who needs Spatial Sampler?
Spatial Sampler is a collection of complete, step-by-step workflows using real data that run comparative tests between different experimental conditions. The worked examples act as a ‘cheat sheet’ for spatial omics analysis. Most existing tutorials stop at the quality control and clustering stages, and a need for testing between groups is what inspired Sarah to develop Spatial Sampler:
When I started working with spatial data, I could find tutorials for how to load, QC and annotate my datasets. But I struggled to find complete workflows for testing experimental questions. I wanted a collection of examples using real data - so that’s what Spatial Sampler aims to provide. I had inspiration from amazing existing resources such as The R Graph Gallery. I’m now looking forward to others using Spatial Sampler and contributing more tools and resources that the community can use.”
Different elements of Spatial sampler will help different people:
Newcomers to spatial omics can apply worked examples to their own datasets
Busy bioinformaticians can use code snippets to run specific tests without rewriting code
Trainers can readily adapt open source instructions for their own training.
What is spatial omics?
Spatial omics combines the power of single-cell transcriptomics with spatial localisation, allowing researchers to measure gene expression and morphology while retaining the individual locations of transcripts within their native tissue environment. By inheriting analytical strengths from both single-cell technologies and microscopy, it enables the testing of complex new hypotheses about expression patterns and cellular interactions that are lost in data from a single modality.
This exciting new field is now moving from a niche to a fully-fledged production analysis technique, and given the rapid growth in utilisation, analysis methods are constantly being invented as new applications are discovered. Now that it is possible to generate high-quality data at scale, the downstream analysis options have expanded accordingly to help answer specific biological questions.
An example of this is in quantifying changes in the tumour microenvironment, such as changes in cell type composition between tumour subtypes, changes in immune cell migration between groups, or expression changes in different cells by experimental group or tissue region.
Spatial Sampler was initially developed by Sarah to analyse data from the Griffith Central Facility Genomics for A/Prof Nic West and Dr Amanda Cox (Griffith University) relating to their research projects in the mucosal immunology and cancer immunology spaces.
Sarah’s role at QCIF Digital Research is co-funded by Australian BioCommons.
Register now for the workshop that will provide a practical introduction to comparative analysis of spatial omics data using Spatial Sampler
Meet the team: Kylie Davies, Senior Business Analyst
Our team members bring deep expertise and broad experience. Hear how a Business Analyst contributes to the BioCommons mission from Kylie Davies.
Describe your role at Australian BioCommons
My role with Australian BioCommons has been to work as a business analyst for the GUARDIANS project - a multi-organisational project aimed at uplifting Australia's Genomic research capabilities and opportunities through the creation of national infrastructure that brings down barriers to the secure sharing of human genomic data. The better and quicker this access can be, the better and broader the opportunities to solve complex and challenging human diseases.
How can Business Analysts improve infrastructure and/or scientific research?
The role of the Business Analyst (BA) is always multifaceted. At the start of a project the BA focuses heavily on unpacking the project work packages with the input of subject matter experts, and translating these into detailed use cases and requirements. Even in Agile units this work is critical. The BA sits between the technical development team and the Project Management (PM) team; and between the future users and the project team (both technical and PM).
“Your role is to get into the weeds enough to unpick the key deliverables and major challenges, while managing the expectations of stakeholders to help the team achieve delivery of a technical product that is fit for purpose and delivers the key requirements.”
A good BA will have experience and some understanding of technical matters, User Experience considerations and project scope. They should be a good listener and good at eliciting information in a relaxed way. We are just one part of the team, but we make a vital contribution in ensuring the planned infrastructure is delivered in such a way that it meets the real needs of users, with all parties being clear about the key deliverables from a business value perspective. Even research infrastructure has a measured business value. If this is delivered, not only does it serve scientific researchers better, but it means the business case for future improvements (and further project funding) easier to make.
Tell us about the impact of your role
I think for some of the project teams across the GUARDIANS project and its predecessor project, on which I also worked, they had not worked through requirements with a Business Analyst before. They found that the process clarified the details of their plans and freed them up to get cracking on development work without circular discussions and going down too many rabbit holes mid-development.
In the research world, it is necessary to allow for enquiry and discussion. It's different from commercial environments in that we need to allow flexibility for exploration. I needed to learn that when I first worked with researchers starting in 2021. There is a necessary tension between maintaining freedom to explore and avoiding paralysis by analysis.
I think probably one impact I had as the BA was to convene forums and exercises for that exploration to really flourish, but then encourage decisions to be made at a certain point through the documentation of requirements. The feedback I have received indicated that the project teams felt that they were clearer about their goals and felt heard, and yet unified, in their development plans as a result of the work we did together. Then as the products were rolled out, they could demonstrate to wider audiences not only what they had developed, but how each element served use cases.
I should add I have learned a lot too in my role from other BAs. There are a couple of other BAs working on GUARDIANS and it's been a pleasure and learning experience particularly to see them work with stakeholders. We also have a highly experienced BA inhouse at BioCommons in the form of Farah Zaib Khan and I am in awe of Farah's work. We have caught up a few times to share techniques.
What’s next?
I am retiring soon. I am happy to be retiring although it's come a little earlier than I had originally planned, due to my own cancer diagnosis mid 2023. I continue to be well but it is a progressive form of disease so the medical interventions are becoming more frequent, but still not too terrible.
I am living proof of the benefits of medical research. The prognosis for the type of cancer I have was extremely poor a decade ago, but I have been able to access three new cancer drugs delivered to us very recently by clinical research. The drug I am on now was TGA approved in 2025! This has delivered tangible and hugely valuable benefits for me. I have been able to work and enjoy a good quality of life. I undertook a wonderful adventure holiday in 2025. Thank you to Australia's research community for this.
2026 will see me nesting more at home. I have a large garden and grow some produce and I am aiming to ramp that up as soon as I finish work. I have adult children who are some of my favourite humans and plan to catch up with them and friends a bit more. And fish and boat with my husband who is retiring at the same time.
I will continue to be involved in two voluntary consumer representative groups for cancer treatment centres, and I hope to follow the progress of GUARDIANS and BioCommons as a remote observer.
Australian BioCommons and Southern eDNA Society partner to deploy national infrastructure
Australian BioCommons and the Southern eDNA Society (SeDNAS) have signed a Memorandum of Understanding (MoU) to acknowledge, and work towards, the shared infrastructure challenges faced by life science researchers performing eDNA analysis.
Australian BioCommons and the Southern eDNA Society (SeDNAS) have signed a Memorandum of Understanding (MoU) to acknowledge, and work towards, the shared infrastructure challenges faced by life science researchers performing eDNA analysis. By collaborating with the community to document the current research landscape, this partnership aims to deploy national-scale bioinformatics infrastructure that supports life science researchers across Australia and New Zealand.
What is eDNA and why is this needed?
Environmental DNA (eDNA) involves the capture of nucleic acids from an environment, such as water, soil or air, to identify the organism in that environment without needing to visually observe them.
This method allows researchers to cost-effectively monitor biodiversity, track invasive pests, and detect rare species over large areas with minimal disturbance to the organisms (such as animals or bacteria) being identified. However, the rapid adoption of these methods across academia, government, and industry has created a pressing need to better coordinate the digital infrastructure and data resources required to support this growing community. eDNA analysis generates massive amounts of data that requires comparison to curated reference databases, such as GOLD (Genomes OnLine Database) and SILVA. As such, practitioners are facing challenges regarding the availability and coordination of the necessary digital infrastructure.
The partnership between SeDNAS and Australian BioCommons
To leverage combined expertise and resources, this partnership brings together the SeDNAS community of experts and practitioners with the bioinformatics infrastructure experience of Australian BioCommons. The collaboration recognises that science benefits from researchers working together, and that shared national infrastructure can help alleviate the identified challenge.
Next steps: building the roadmap
Key activities to support the eDNA community include:
Community consultation engagement activities, such as surveys and community meetings, to collect the specific bioinformatics infrastructure requirements of life science researchers
Strategic planning to develop targeted, community-endorsed infrastructure roadmaps that describe the national bioinformatics research infrastructure needed to support the field
Capacity building by raising awareness of existing infrastructure and collaborating on priorities such as training, upskilling, and data resource development.
Get involved
Empowering wet-lab postgraduate students with computational and communication skills
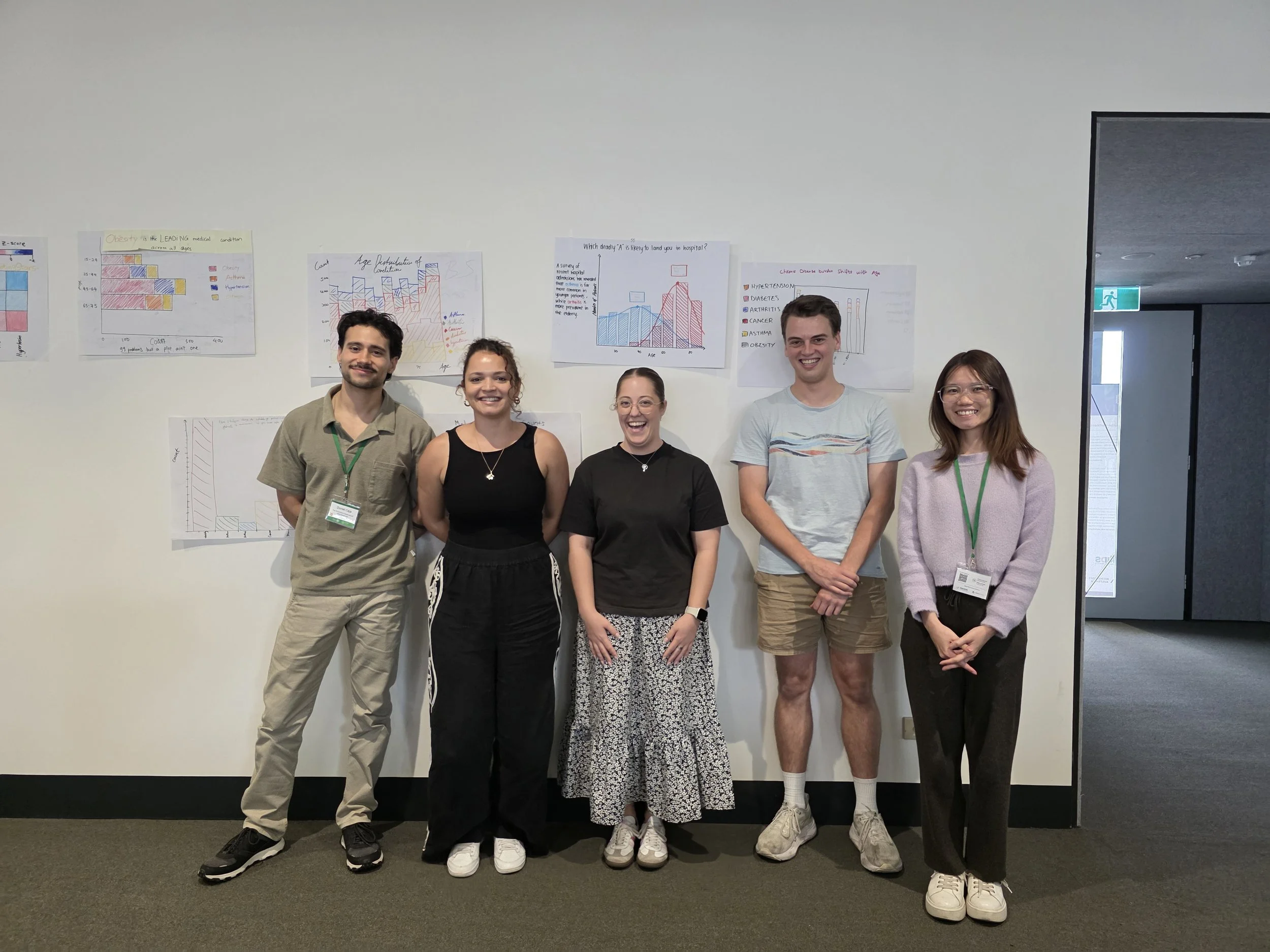
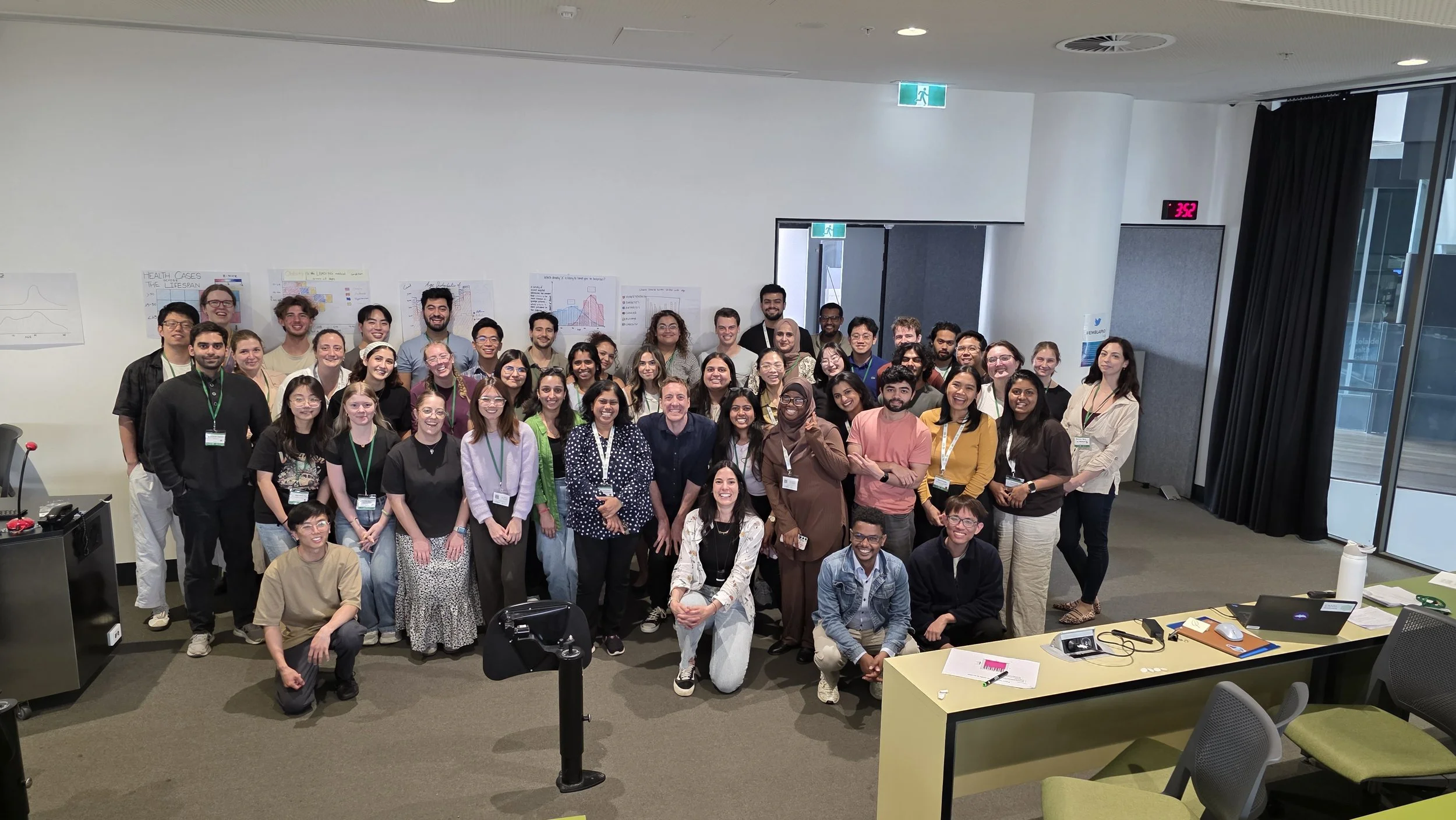
Australian BioCommons team members Dr Giorgia Mori and Dr Benjamin Goudey delivered a session on data visualisation and AI/ML tools as part of EMBL Australia’s 10th annual PhD course.
Australian BioCommons team members have helped to equip the next generation of researchers with the skills and confidence to work at the cutting edge of data-driven science.
As part of EMBL Australia’s 10th annual PhD course, consisting of over 60 high-achieving PhD students in the early part of their research careers, Dr Benjamin Goudey and Dr Giorgia Mori joined other experts at South Australian immunoGENomics Cancer Institute (SAiGENCI) to deliver sessions across the two-week program.
The topics ranged from emerging life sciences technologies to Artificial Intelligence (AI) foundations and scientific modelling, and also included science communication and ethics. The goal of BioCommons’ involvement was to help bridge the gap between the wet-lab results and dry-lab computational skills that make up modern scientific research.
Dr Giorgia Mori’s training expertise had the room buzzing as students worked in small groups to identify the most effective data visualisation methods to explore how to tell a compelling story with their data.
BioCommons’ AI Technical Lead, Dr Benjamin Goudey, delivered a dynamic crash course regarding these topics, giving a clear, high-level understanding of when and why AI/ML can be an appropriate tool in the life sciences.
As noted in EMBL Australia’s event report, Madison Hindes from Adelaide University came away with a new appreciation for dry-lab science:
“I definitely thought science happened in a lab until coming here and now I can appreciate there is much more to it than that.”
Our dedication to diversity, disability and inclusion
BioCommons’ new Diversity, Disability and Inclusion position statement reflects our commitment to provide a safe, fair and enriching environment for all our staff, collaborators and broader community. It forms part of our continuous journey towards creating a workplace where everyone belongs.
Australian BioCommons aspires to be a project where all people are valued, respected, and supported to realise their full potential. Our new Diversity, Disability and Inclusion position statement reflects our commitment to turn that aspiration into action and provide a safe, fair and enriching environment for all our staff, collaborators and broader community, one that reflects the evolving expectations of the diverse partners and sectors we serve.
Our dedication to diversity, disability and inclusion is more than just a statement, it’s a continuous journey towards creating a workplace where everyone belongs. How we approach inclusion was discussed with the whole Hub team at our recent Offsite meeting, where we established our inaugural Inclusion Committee. Our commitment has been added to the About section of our website where we describe what we do and how we do it, alongside our existing Code of Conduct. Both of these documents embody the BioCommons values of Respect, Integrity, Collaboration, Communication, and Innovation.
By fostering a culture of accountability to our colleagues, collaborators, and the communities we serve, Australian BioCommons is committed to driving meaningful change within our team and across the broader scientific community.
Australian BioCommons Hub Team
Solving the big international bioinformatics challenges from the comfort of home
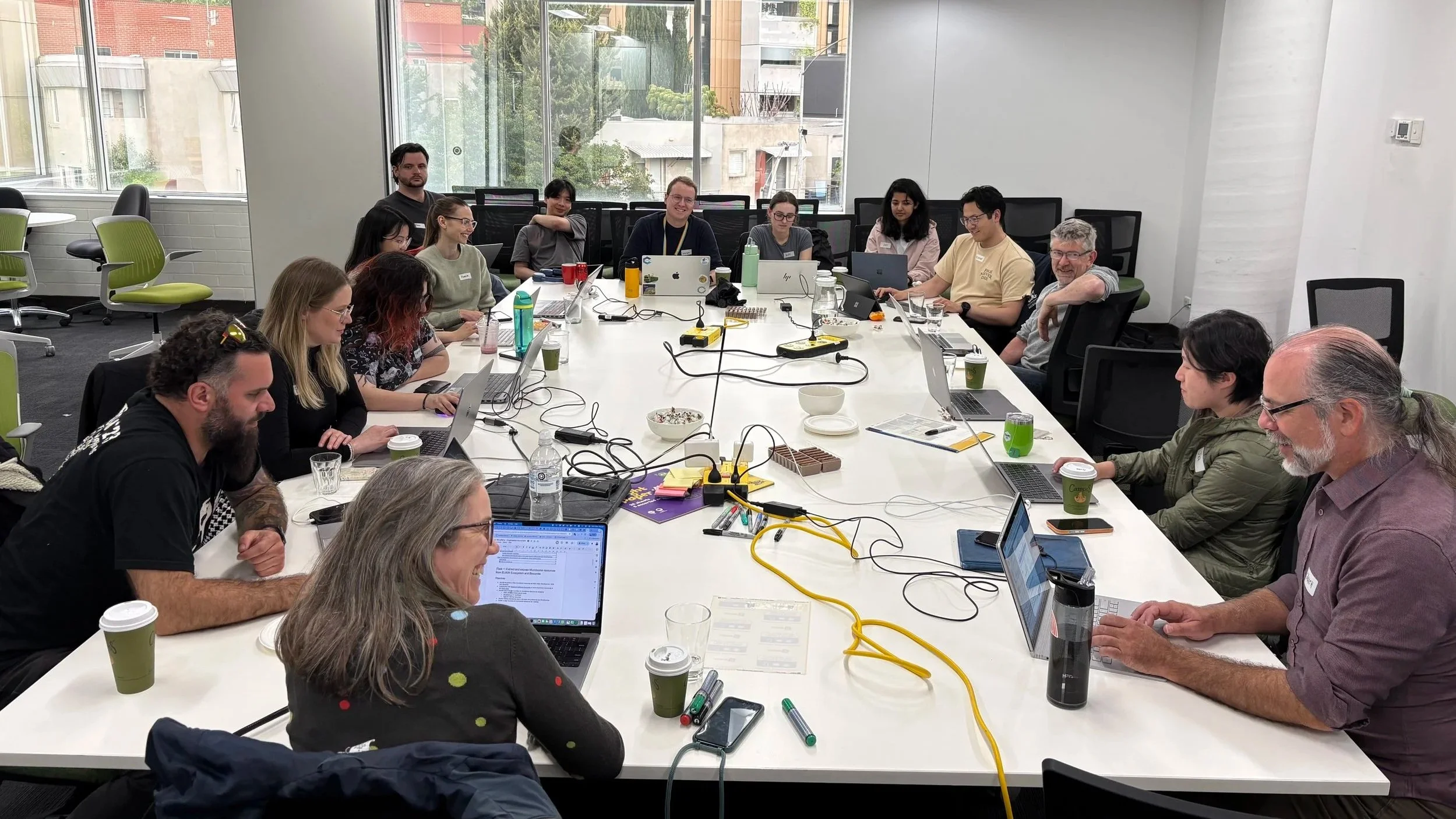
The Australian Outpost of ELIXIR’s BioHackathon Europe, running for the fourth consecutive year, recently brought together 16 bioinformaticians from across Australia to the University of Melbourne for an intensive week of collaborative coding.
The Australian Outpost of ELIXIR’s BioHackathon Europe, running for the fourth consecutive year, recently brought together 16 bioinformaticians from across Australia to the University of Melbourne for an intensive week of collaborative coding.
Facilitated by BioCommons’ ongoing partnership with ELIXIR, the event provided a unique opportunity for local experts to contribute directly to global bioinformatics projects, connecting with their local peers and linking into the main BioHackathon Hub in Berlin.
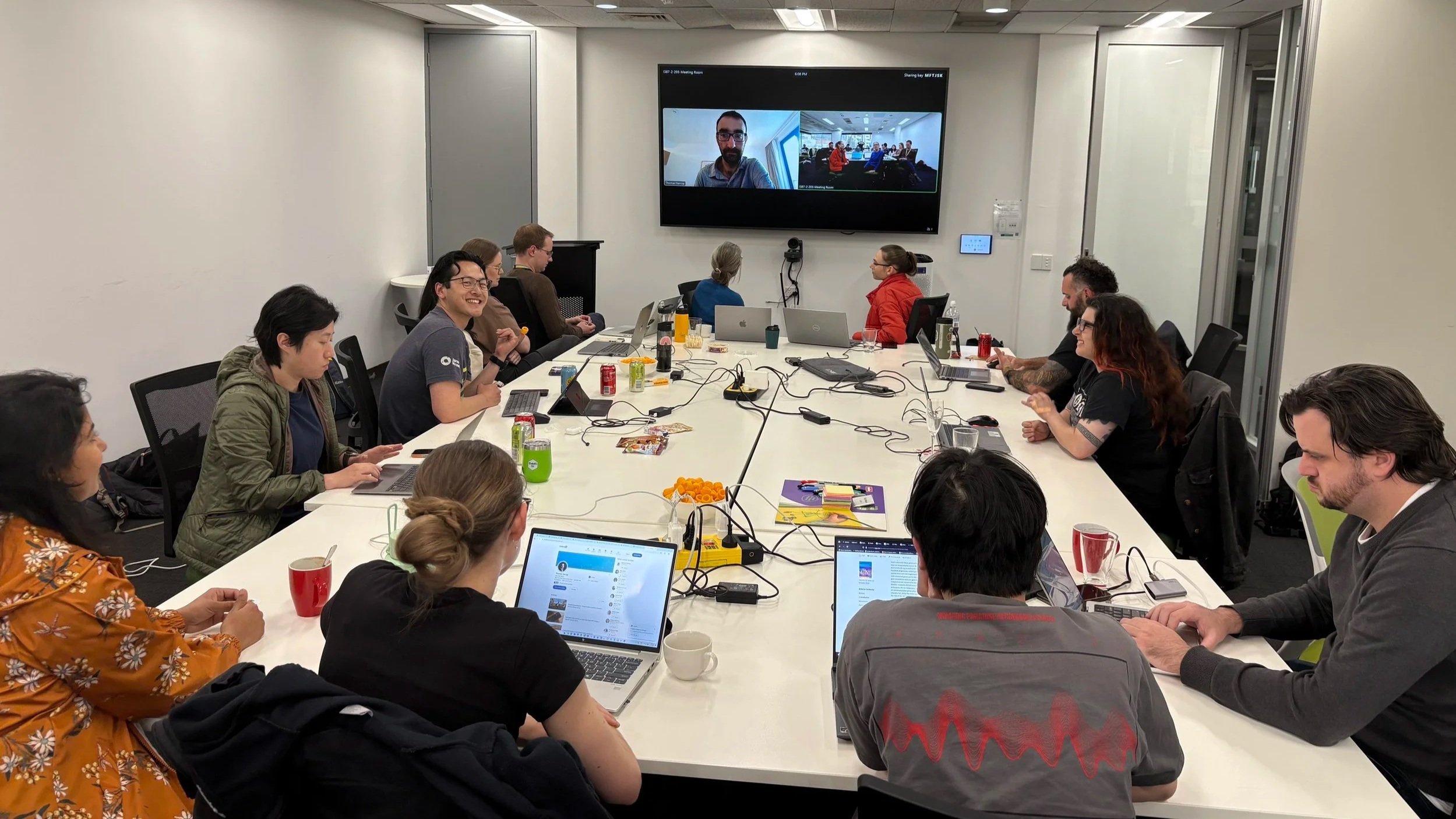
From 3-7 November, the Australian team worked from 12:00 - 8:30 PM AEDT each day to ensure overlap with their European counterparts. This year’s outpost dedicated its efforts to three key projects.
Team members collaborating during the BioHackathon
One team tackled the challenge of using BUSCO (Benchmarking Universal Single-Copy Orthologs) tools for phylogenomics. BUSCO genes are standard for assessing genome assembly and annotation completeness, but their use in phylogenetics is often challenged by gene duplication (paralogy). Representatives from BioCommons, University of Melbourne, University of Sydney, Curtin University, and University of Tasmania collaborated with peers around the world to build an automatic pipeline that explicitly resolves these paralogy events, which will result in larger, more informative datasets for biodiversity research.
Another team brought together experts from BioCommons, Curtin University, University of Melbourne, QIMR Berghofer, and University of Queensland to focus on expanding the benchmarking landscape for long-read transcriptomics. While foundational benchmarks for long-read RNA-Seq were established in 2020, the field has evolved rapidly since then with new tools and protocols. This project worked towards an updated benchmark, leveraging new datasets to ensure the reproducibility and comparability of modern transcriptomics research.
The final project brought together a ‘supergroup’ from the Australian Tree of Life (ATOL) and European Reference Genome Atlas (ERGA) projects, to tackle streamlining FAIR metadata for biodiversity genome annotations. Tom Harrop represented the Outpost in-person in Berlin, with Tiff Nelson co-leading the whole project from the Australian Outpost, along with participants from BioCommons, Melbourne Bioinformatics, University of Queensland, and QCIF. With thousands of reference genomes being generated globally, there is an urgent need for structured, automated ways to report metadata for genome annotations. This project worked on developing automated tools to process, validate, and deploy this metadata, ensuring it is harmonised and discoverable across global repositories.
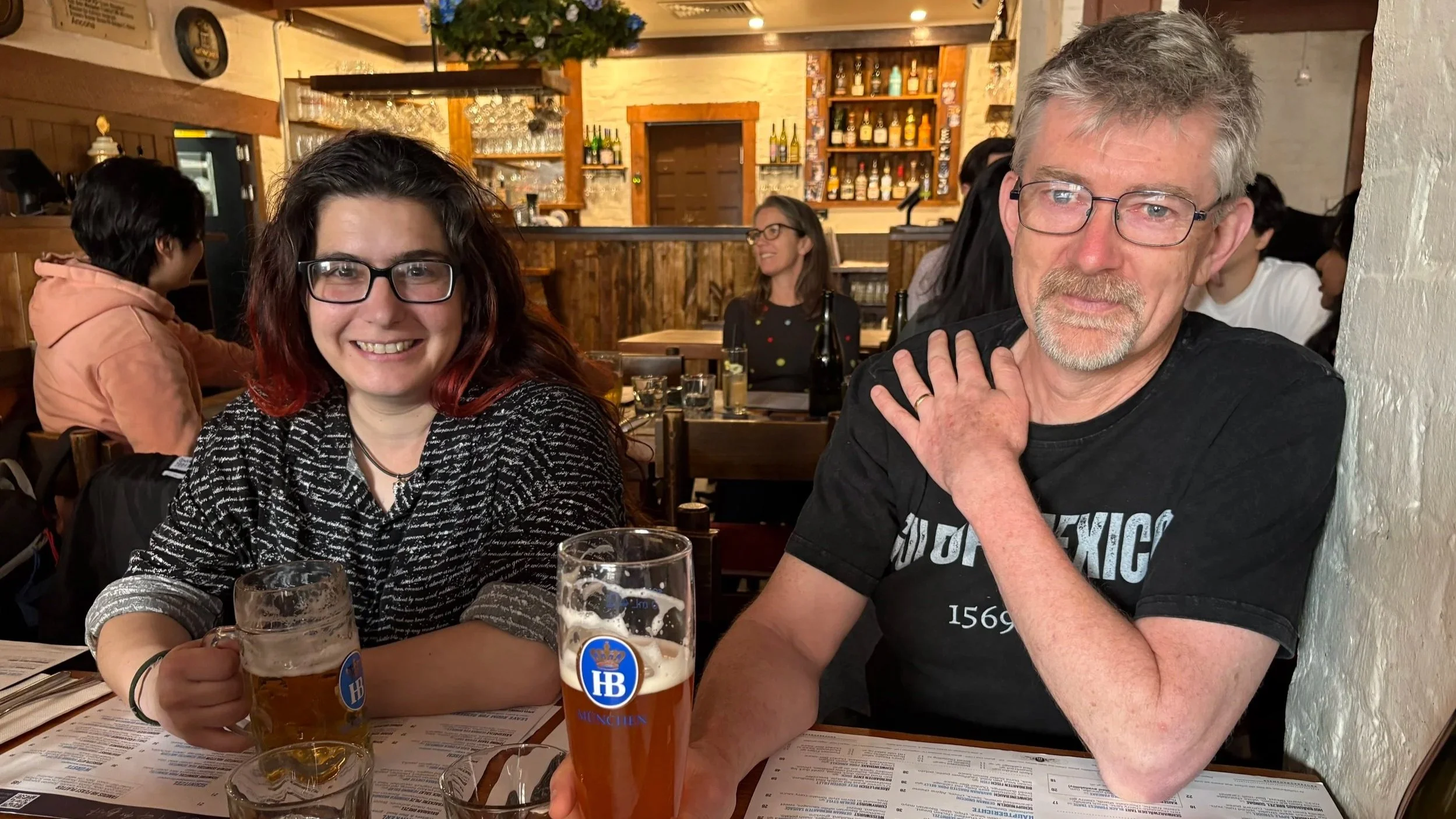
Outside of the intensive project work, the local team built their professional relationships over lots of Melbourne’s coffee and terrible weather. Hacking paused for the ‘race that stops the nation’ (for approximately four minutes!) on Melbourne Cup day, and the event wrapped up with a celebratory German lunch at the iconic Hofbräuhaus Melbourne to feel more connected to their colleagues at the main site of the BioHackathon Europe.
“This was a fantastic event. It's rare to get the opportunity to work as part of a specialised software development team while in academic research. This was a great opportunity to learn the skills required to make my code more usable, work together to produce larger projects, and plan contributions amongst team members,” said Thomas Crow, University of Queensland PhD candidate and affiliate of the ARC Centre of Excellence for Plant Success in Nature and Agriculture.
“The event covered important pipeline and repo development skills I have been seeking for some time. That our collective work has also led to collaborations in Europe, potential publications and research impact make this event even more worthwhile. I am very excited to attend future events."
The collaboration continues, and we look forward to bringing another team together for BioHackathon Europe 2026. Be sure to watch for our call for applications in mid-2026 if you'd like to participate!
Read ELIXIR’s article about the event: Accelerating open science - the impact of the BioHackathon Europe
BioHackathon attendees avoiding the Melbourne weather
Celebratory German lunch at the iconic Hofbräuhaus Melbourne
Dialling in for discussions with Tom Harrop in Berlin
National supercomputing and expertise to enable the Australian Fish Genomics Initiative
The Australian Fish Genomics Initiative (AFGI) is generating high-quality reference genomes for 85 species to improve population monitoring and environmental assessment, and support sustainable fisheries and aquatic conservation.
Fish populations across Australian freshwater and marine ecosystems are struggling to cope with the combined threats of invasive species, pollution, and climate events. A significant proportion of Australia’s 5,000 fish species remains poorly understood, lacking the essential referential biomolecular data that could inform improved population monitoring, environmental assessment, and the sustainable management of fisheries and aquatic ecosystems. This conservation issue is exemplified by the critical gap in genomic data, with only 3% of marine species and 25% of freshwater species even having species-level data.
The Australian Fish Genomics Initiative (AFGI) is responding to this dire situation by characterising 85 species of Australia’s fish species through the generation of high-quality reference genomes.
Queensland groper (Epinephelus lanceolatus). Image credit: Getty images
Minderoo Foundation and Bioplatforms Australia established the Initiative with the aim to collaborate nationally with marine and freshwater research communities to generate essential data for researchers and practitioners to enhance our understanding of species adaptation, population dynamics, and evolutionary relationships.
The AFGI consists of 56 partner projects that will generate genomics, genetics (e.g., population genetics) and transcriptomics data from fish species from all major Australian habitats: freshwater, estuarine and marine environments. Analysing this huge volume of high-quality data presents a challenge that fish biologist and AFGI Bioinformatician, Dr Amy Tims, has been tasked with solving. Amy joined the BioCommons team to facilitate AFGI’s access to national supercomputing resources and specialised bioinformatics expertise. Data is already arriving via the Bioplatforms Australia Data Portal, and Amy is providing hands-on assistance to consortium partners to perform data QC, genome assembly, and data curation.
"We need to look at population connectivity to manage fisheries sustainably, and understand genetic diversity to implement effective conservation breeding programs,” said Dr Tims.
“We also need to know what is happening at the genetic level to determine whether a species can survive extreme weather conditions. In short: we need genomes!"
A dedicated compute allocation at Pawsey Supercomputing Research Centre (Pawsey) has been provided to all AFGI partners via the Australian BioCommons Leadership Share (ABLeS), removing the computational constraint that can be a barrier to entry for researchers. This is complemented by the Australian Tree of Life (AToL) project that is building systems to automate the complex process of genome assembly and publication at scale. This combination of dedicated expertise and national infrastructure is already delivering results, and the first draft genomes have already been assembled.
The high quality data that AFGI is generating will unlock new insights into fish biodiversity, evolution, conservation, aquaculture, and biosecurity, including pest and disease management. It demonstrates that strategic collaboration and the right research infrastructure can support critical research that addresses our biggest challenges, like the sustainable management of Australia’s aquatic ecosystems.
GUARDIANS partners gather in Brisbane to align on national human omics infrastructure
Partners collaborating on the future of Australia’s human omics research data ecosystem gathered in Brisbane this month for their second in-person meeting. This effort is driven by the GUARDIANS program that is delivering digital research infrastructure for human omics data nationally.
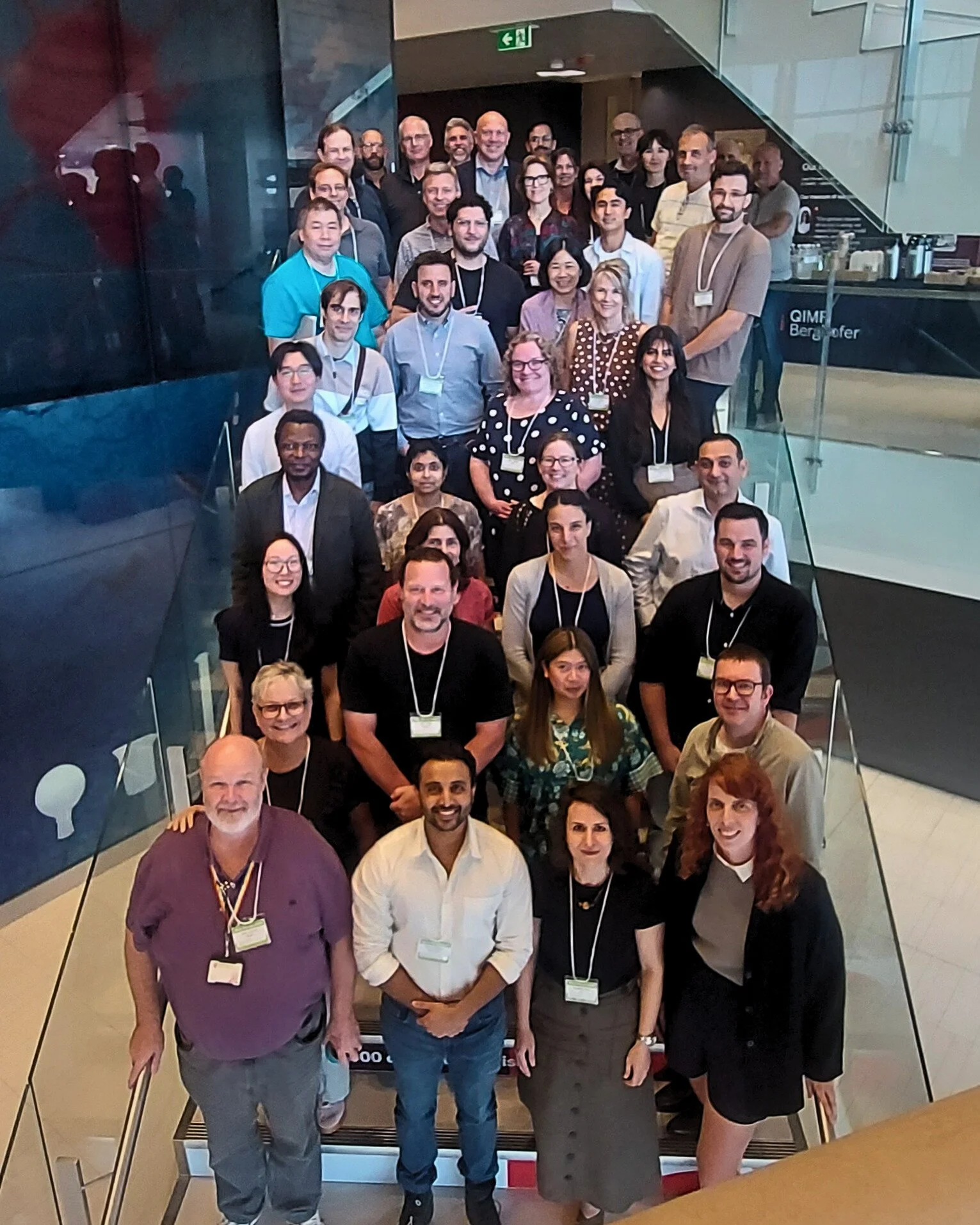
Partners collaborating on the future of Australia’s human omics research data ecosystem gathered in Brisbane this month for their second in-person meeting. This effort is driven by the GUARDIANS program that is delivering digital research infrastructure for human omics data nationally.
Building a complex national ecosystem for this data demands more than just the technical tools, it needs active collaboration and a genuine community of practice. Coming together to review the foundational achievements of Year 1 of the program and strategically align activities for Year 2, the meeting fostered an environment of open communication, mutual learning, and collective problem-solving across the two days.
The first day of the program focused on partner presentations, with teams highlighting their progress and challenges. These covered data commons deployments, data access control frameworks, and platform integration, and the presentations concluded with an engaging retrospective discussion.
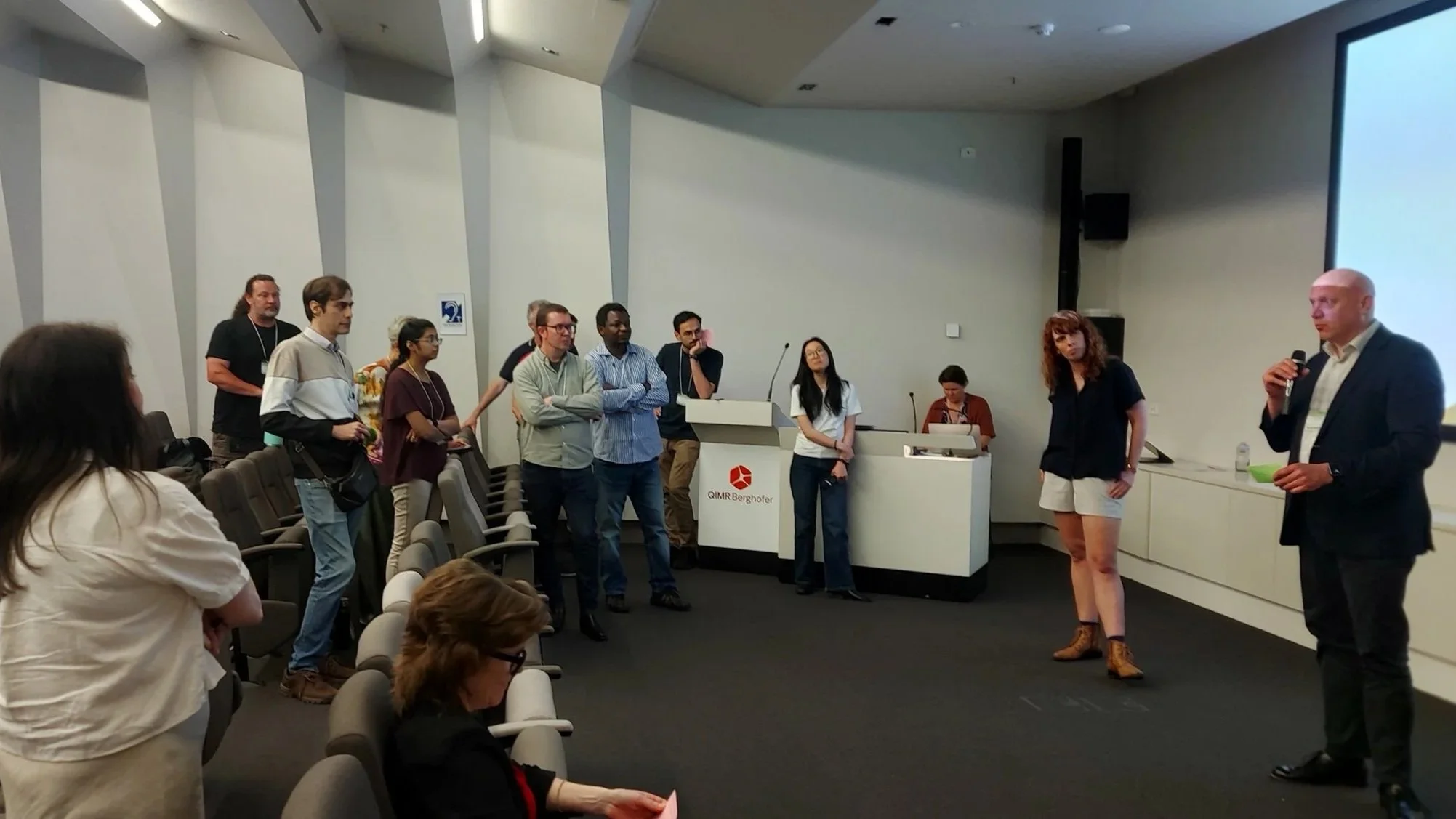
A/Prof Natalie Taylor, UNSW, speaking on ‘Implementation to Impact’
Partners and GUARDIANS team members at QIMR Berghofer
A highlight of the day was the keynote presentation on ‘Implementation to Impact’ from A/Prof Natalie Taylor, UNSW, speaking to systematic approaches to bridge the gap between research evidence and clinical practice. She challenged the attendees to design infrastructure not just for technical functions, but for real-world adoption to deliver real health outcomes.
The second day focused on practical applications, featuring a series of workshops to showcase the progress and frameworks across the project. An integrated data access and release management workflow was demonstrated by the Collaborative Centre for Genomic Cancer Medicine, the Garvan Institute of Medical Research showcased their Elsa, CTRL and REMS tools, and QIMR Berghofer gave a highly practical review of consent and the data access request process.
The session from the GUARDIANS’s Ethical, Legal, and Social Implications (ELSI) team navigated through the complexities of cross-border omics data governance. This was complemented by the ‘Threat Modelling’ workshop run by the BioCommons Cybersecurity Specialist. Finally, GUARDIANS Project Managers led a workshop to identify concrete opportunities for inter-organisational interoperability in Year 2, ensuring future efforts are integrated rather than siloed.
The strategic discussions and practical demonstrations in Brisbane showed that progress is being made towards a national human omics research data ecosystem.
“Reflecting on Year 1, the significant outcomes include technical tools and infrastructure, and also the genuine community of practice and collaborative environment we’ve built,” said Prof Bernie Pope, A/Director (Human Genome Informatics) at BioCommons and Lead for the GUARDIANS project.
“This foundation will be crucial as we tackle increasingly complex challenges in delivering this national data ecosystem. ”
ELSI presentation on ‘Omics Data Beyond Borders - Law, Ethics and Governance’
GUARDIANS team and partners discussing collaboration opportunities
The GUARDIANS program is accelerating human omics research in Australia through the development of world-class digital infrastructure. The program is led by Australian BioCommons with contributions from partner organisations including the Australian Access Federation, Children’s Cancer Institute Australia, Garvan Institute of Medical Research, National Computational Infrastructure, QIMR Berghofer Medical Research Institute, The University of Melbourne, and The University of Sydney. GUARDIANS forms part of Australian BioCommons’ Human Genome Informatics Initiative and receives National Collaborative Research Infrastructure Strategy (NCRIS) support through Bioplatforms Australia.
AI for Science Australian Hackathon
Could you benefit from some free expert advice to accelerate your research? The AI for Science Hackathon is an exciting opportunity for Australian life science projects to receive support to realise performance gains and speedups in their computation.
Could you benefit from some free expert advice to accelerate your research? The AI for Science Hackathon is an exciting opportunity for Australian life science projects to receive support to realise performance gains and speedups in their computation.
In collaboration with NVIDIA and OpenACC organization, BioCommons has partnered with NCI, Monash University, Pawsey Supercomputing Research Centre and SHARON.AI to support an in-person hackathon in Melbourne next February.
Teams will be paired with experienced computational mentors based on their projects, bringing together the programming models, libraries, tools or specialised compute resources to port, accelerate and optimise their applications on whichever data centre architecture is relevant to the project.
This multi-day hands-on event is ideal for scientists, researchers and developers working in Australian research institutions, NCRIS facilities, national science agencies and research centres who can bring a team to work on their computational science challenge. Maybe you have some custom code and you want help to apply GPU optimisations? Or you are using an Alphafold workflow that requires tweaking but you need support to optimise the code? Perhaps you’ve developed an AI/ML pipeline that needs refinement? By collaborating with each other and the mentors, participants will leave with their scientific applications accelerated and/or optimised on high performance computers, or at least with a clear understanding of how to leverage these resources.
Applications are now open and BioCommons will be represented on the review committee, so we suggest you discuss the potential of your life sciences project with benjamin@biocommons.org.au asap. To put your project forward for consideration, you must have at least 3 team members and be fluent with the code you are bringing along. Registration is free, but you will need to cover your travel expenses.
Apply now before the 6 Jan 2026 deadline
Australian health research infrastructures take centre stage in Europe
A delegation from Australian BioCommons, Bioplatforms Australia and the NCRIS Health Group recently concluded a series of strategic meetings, strengthening global ties and aligning national strategies with our European counterparts.
A delegation representing Australian health research infrastructures recently concluded a series of strategic meetings in Europe. Australian BioCommons, Bioplatforms Australia and the NCRIS Health Group strengthened global ties and shared insights into shared national priorities with a range of strategic counterparts.
The centerpiece of the trip was attendance at the Australian and European Health and Life Sciences Research Infrastructure Symposium in Prato, Italy. Through a series of facilitated sessions, participants discussed key themes including AI in health and life sciences, workforce development, and the role of research infrastructures as strategic national assets. Set against the backdrop of evolving research policies in both continents, including preparations for the EU’s Framework Programme 10 and the next Australia National Research Infrastructure roadmap, these discussions will help shape a joint position on the value of structured international collaboration and the targeted investment required, and hopefully influence future roadmaps.
Attendees of the Australian and European Health and Life Sciences Research Infrastructure Symposium in Prato, Italy
Building on these high-level strategic discussions, a series of targeted visits demonstrated how these key themes are being put into practice at world-leading institutions. At EMBL Barcelona, Head Dr James Sharpe welcomed the delegation to explore the cutting-edge imaging and microfabrication facilities. Discussions continued at EMBL-EBI, led by Interim Director Jo McEntyre, and focusing on AI implementation, data archiving and spatial omics, setting the stage for her upcoming visit to Australia in November to continue these collaborative talks.
EMBL Barcelona and EMBL-European Bionformatics Institute showcased their facilities, projects and perspectives. In the United Kingdom, the team met with leaders of the Darwin Tree of Life project at the Wellcome Sanger Institute. These meetings were helpful in aligning the BioCommons’ Australian Tree of Life project with its international peer, and for holding further discussions around the integration of AI into the scientific process across the domains of data access and training. Finally, there was an opportunity to connect and share insights with the newly formed BioFAIR UK, a like-minded organisation set up to address similar challenges in building a national data, methods and people ‘commons’.
The group included Australian BioCommons Director, Dr Jeff Christiansen, and A/Director, BioCloud, Dr Steven Manos, who joined leaders from Bioplatforms Australia, EMBL Australia, Phenomics Australia, the National Imaging Facility, Microscopy Australia, the Population Health Research Network and Therapeutic Innovation. Reflecting on the visit, Dr Jeff Christiansen said, “It was valuable to have time to reflect on areas of mutual interest, including the application of AI in the life sciences and the critical role that high-quality data plays in underpinning AI approaches. Hearing about new developments at EMBL-EBI to support the long-term preservation of spatial omics data – an exciting intersection of imaging and omics – was particularly informative.”
Fostering an integrated global research ecosystem will ultimately benefit the Australian research community, with opportunities for collaboration around shared priorities.
For another perspective, see the article written by EMBL Australia about the trip.